Abstract
Aims/hypothesis
The mitochondrial chaperonin heat shock protein (HSP) 60 is indispensable in protein folding and the mitochondrial stress response; however, its role in nutrient metabolism remains uncertain. This study investigated the role of HSP60 in diet-induced non-alcoholic fatty liver disease (NAFLD).
Methods
We studied human biopsies from individuals with NAFLD, murine high-fat-diet (HFD; a diet with 60% energy from fat)-induced obesity (DIO), transgenic (Tg) mice overexpressing Hsp60 (Hsp60-Tg), and human HepG2 cells transfected with HSP60 cDNA or with HSP60 siRNA. Histomorphometry was used to assess hepatic steatosis, biochemistry kits were used to measure insulin resistance and glucose tolerance, and an automated home cage phenotyping system was used to assess energy expenditure. Body fat was assessed using MRI. Macrophage infiltration, the lipid oxidation marker 4-hydroxy-2-nonenal (4-HNE) and the oxidative damage marker 8-hydroxy-2′-deoxyguanosine (8-OHdG) were detected using immunohistochemistry. Intracellular lipid droplets were evaluated by Nile red staining. Expression of HSP60, and markers of lipogenesis and fatty acid oxidation were quantified using RT-PCR and immunoblotting. Investigations were analysed using the two-way ANOVA test.
Results
Decreased HSP60 expression correlated with severe steatosis in human NAFLD biopsies and murine DIO. Hsp60-Tg mice developed less body fat, had reduced serum triglyceride levels, lower levels of insulin resistance and higher serum adiponectin levels than wild-type mice upon HFD feeding. Respiratory quotient profile indicated that fat in Hsp60-Tg mice may be metabolised to meet energy demands. Hsp60-Tg mice showed amelioration of HFD-mediated hepatic steatosis, M1/M2 macrophage dysregulation, and 4-HNE and 8-OHdG overproduction. Forced HSP60 expression reduced the mitochondrial unfolded protein response, while preserving mitochondrial respiratory complex activity and enhancing fatty acid oxidation. Furthermore, HSP60 knockdown enhanced intracellular lipid formation and loss of sirtuin 3 (SIRT3) signalling in HepG2 cells upon incubation with palmitic acid (PA). Forced HSP60 expression improved SIRT3 signalling and repressed PA-mediated intracellular lipid formation. SIRT3 inhibition compromised HSP60-induced promotion of AMP-activated protein kinase (AMPK) phosphorylation and peroxisome proliferator-activated receptor α (PPARα levels), while also decreasing levels of fatty acid oxidation markers.
Conclusion/interpretation
Mitochondrial HSP60 promotes fatty acid oxidation while repressing mitochondrial stress and inflammation to ameliorate the development of NAFLD by preserving SIRT3 signalling. This study reveals the hepatoprotective effects of HSP60 and indicates that HSP60 could play a fundamental role in the development of therapeutics for NAFLD or type 2 diabetes.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
Non-alcoholic fatty liver disease (NAFLD) is a public health issue and a major risk factor for cardiovascular disease, type 2 diabetes and chronic kidney disease [1]. Mitochondrial dysfunction accelerates the development of NAFLD; indeed, a growing number of studies have revealed that mitochondrial dysfunction downregulates hepatic lipid homeostasis and enhances reactive oxygen species (ROS) production to aggravate lipid peroxidation, inflammatory cytokine production, apoptosis, and fibrosis in hepatic tissue [2,3,4].
Mitochondrial heat shock protein (HSP)60 plays an important role in controlling mitochondrial stress [5]. This chaperonin is also present in the cytosol, the cell membrane and body fluid, being involved in regulating inflammation and cardiometabolic and neurodegenerative diseases [6, 7]. Loss of HSP60 has been associated with high-fat-diet (HFD)-induced obesity (DIO) and diabetes in mice [8]. More specifically, HSP60 interference has been shown to cause mitochondrial dysfunction and insulin resistance [8], whereas HSP60 upregulation by metformin attenuates diabetes-induced mitochondrial dysfunction [9]. Rosiglitazone, an early insulin sensitiser (previously used to treat diabetes), reverses obesity-induced mitochondrial protein loss in ob/ob mice. Moreover, palmitate oxidation and HSP60 protein levels are promoted in adipocytes from rosiglitazone-treated mice [10]. Despite these investigations, the specific biological role that this chaperonin may play in the development of NAFLD remains unclear.
The sirtuin family (consisting of seven members) is comprised of highly conserved class III histone deacetylases that reside in mammalian cells [11]. Sirtuin 3 (SIRT3) is present in the mitochondrial compartment, where it removes the acetyl group from mitochondrial proteins and regulates energy metabolism and oxidative stress [12, 13]. SIRT3 catalyses HSP10 deacetylation and modulates the biological activity of the HSP60/HSP10 chaperone complex, which enhances the folding of the fatty acid oxidation enzyme medium-chain acyl-CoA dehydrogenase [14]. Sirt3 knockout mice develop severe hepatic steatosis upon HFD consumption [15], while Sirt3 overexpression promotes lipophagy, and represses lipotoxicity in hepatocytes incubated in palmitic acid (PA) [16]. Interestingly, muscle- and liver-specific Sirt3 knockout mice did not manifest any overt metabolic phenotype following HFD consumption, despite a marked global hyperacetylation of mitochondrial proteins [17]. This observation suggests that a complex coordinated defect may exist in Sirt3 in multiple tissues in the germline Sirt3 knockout mice.
This study aimed to utilise human specimens from individuals with NAFLD and Hsp60 transgenic (Tg) mice to investigate whether HSP60 affects lipid metabolism during the development of HFD-mediated NAFLD and to elucidate the role of HSP60 in regulating lipogenesis in hepatocytes.
Methods
Cell cultures
Human HepG2 cells (American Type Culture Collection, Manassas, VA, USA) were incubated in DMEM with 10% (vol./vol.) FBS, antibiotics (100 IU/ml penicillin and 100 μg/ml streptomycin), and l-glutamine in a 37°C and 5% CO2 humidified incubator. There was no mycoplasma contamination in HepG2 cells. See the Human Cell Line DNA Typing Report in the electronic supplementary material (ESM) for details.
HFD feeding
All animal studies were approved by the Institute of Animal Care and Use Committee (IACUC) of Kaohsiung Chang Gung Memorial Hospital, Taiwan (no. 2016122605). Mice overexpressing HSP60 driven by mitochondrial phosphoglycerate kinase 1 (FVB/N-Tg[PGK-HSPD1]Wfs/Narl; strain no. RMRC13343; www.nlac.narl.org.tw/RMRC/webc/html/data/show.aspx?ix=1&page=1&kw=Hsp60, accessed 22 January 2018) obtained from the Rodent Model Resource Center, National Laboratory Animal Center, Taiwan) were bred and genotyped as previously described [18]. Animals were group-housed in individual ventilation cages in a specific pathogen-free vivarium (Center for Laboratory Animals, Kaohsiung Chang Gung Memorial Hospital, Taiwan) with a 12 h/12 h light/dark cycle and high-efficiency particulate air conditions (cabinet temperature, 22 ± 2°C and relative humidity, 70 ± 5%). Animals were provided with sterile wood chip bedding and given drinking water and feed ad libitum. Male 8-week-old mice were included and randomised into chow diet (CD)- or HFD-feeding groups. Wild-type (WT) mice (C57BL/6JNarl or FVB; National Laboratory Animal Center, Taipei, Taiwan) or Hsp60-Tg mice were fed an HFD (a diet with 60% energy from fat; catalogue no. D12331; LabDiet, Louis, MO, USA) or standard CD (10% energy from fat; catalogue no. D12329; LabDiet) for 6 months. The HFD and CD diets contained ~20% protein and their mineral and vitamin content met the requirements of the American Institute of Nutrition for mice [19].
In vivo metabolic investigations
Physical activity (locomotion and rearing) and metabolic profiles were characterised in animals using the Automated Home Cage Phenotyping System (Phenomaster; TSE Systems, Homburg, Germany), according to the manufacturer’s instructions. Mice were accommodated in metabolic cages of a climate chamber (23°C and 12 h dark/12 h light cycle) (TSE Systems) for 2 days, with feed and water ad libitum. Upon acclimation, a high throughput physiological signalling calculation unit (TSE Systems) was used to measure volumes of O2 consumption (\( \dot{V}{\mathrm{O}}_2 \)) and CO2 production (\( \dot{V}{\mathrm{CO}}_2 \)), together with the respiratory exchange ratio (RER) and mouse activity over 24 h.
MRI
Mice were anesthetised using isoflurane and MRI was conducted using a 9.4T MRI scanner (Biospec 94/20; Bruker, Ettlingen, Germany). In brief, whole-body images were scanned using a 112/72 mm transmit-receive volume coil. Multi-slice turbo rapid acquisition together with refocused echoes sequence (repetition time/echo time, 3000 ms/9 ms; resolution, 182 × 182 × 2000 μm; 30 slices and number of excitations (NEX)/averages, 8) was utilised to visualise the MRI images. The volume of adipose tissue in the images was calculated using semiautomatic image analysis software (ParaVision PV5.1 software; Bruker).
Measurement of IPGTT and IPITT
Glucose tolerance and insulin tolerance were investigated in mice fed with an HFD for 6 months using IPGTT and IPITT tests, as previously described [20].
Serum biochemistry analysis
Levels of NEFA (Free Fatty Acid Fluorometric Assay; catalogue no. 700310; Cayman Chemical, Ann Arbor, MI, USA), triglyceride (Triglyceride Colorimetric Assay, catalogue no. 10010303; Cayman Chemical), adiponectin (catalogue no. ab108785; Abcam, Cambridge, MA, USA), leptin (catalogue no. ab199082, Abcam) and IL-6 (catalogue no. 431307; BioLegend, San Diego, CA, USA) in sera were quantified using biochemistry or ELISA kits according to the manufacturers’ instructions.
Clinical specimen collection
Studies on human specimens, retrieved from 22 normal liver and 22 fatty liver samples that were archived in the Department of Pathology, Kaohsiung Chang Gung Memorial Hospital, Kaohsiung, Taiwan, were approved by the Chang Gung Medical Foundation Institutional Review Board (202000208B0). HSP60 levels were quantified using immunohistochemistry or immunoblotting (see below). Analysis of protein expression was conducted by individuals who did not know the degree of hepatic steatosis of the specimen. Upon haematoxylin and eosin staining, histopathology was used to analyse fat deposits in the liver, by a pathologist. The area of fat deposition in liver specimens was measured and categorised into four grades: minimal (<5%); mild (5–30%); moderate (31–60%); and severe (>60%) [21].
Immunohistochemistry analysis and scoring
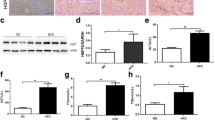
Upon deparaffinisation, blocking by 3% H2O2, and antigen retrieval, sections were incubated in antibodies for HSP60, the oxidative DNA damage marker 8-hydroxy-2-deoxyguanosine (8-OHdG; catalogue no. bs-1278R; Bioss, Boston, MA, USA), the lipid oxidation marker 4-hydroxy-2-nonenal (4-HNE; catalogue no. MAB3249; R&D Systems, Minneapolis, MN, USA), or antibodies for macrophage markers F4/80 (catalogue no. 70076; Cell Signaling Technology, Danvers, MA, USA), integrin αX (also known as CD11c; catalogue no. sc-46676; Santa Cruz Biotechnology, Santa Cruz, CA, USA) or CD206 (catalogue no. sc-58986; Santa Cruz Biotechnology) (see Table 1 for details). Immunosignals were visualised using the Ultravision Quanto Detection System HRP DAB Kit (catalogue no. TL-060-QHL; Thermo Scientific, Waltham, MA, USA) and counterstained using Gill’s haematoxylin. The intensity of immunostained liver tissue was scored as: 0, none; 1, weak; 2, intermediate; and 3, strong. Ten random fields of each section were selected. Immunostained cells were evaluated using a light microscope (Leica, Morrisville, NC, USA). The percentage of immunostained cells in ten random fields (×200 magnification) was calculated. The percentage of 4-HNE+ areas was calculated as mean absorbance by Image J software (Image J 1.51; National Institutes of Health, Bethesda, MD, USA). Macrophages immunostained for CD11c or CD206 in adipocytes were quantified as described in the ESM Methods (Quantification of immunostained cells section).
Assessment of ROS production in liver specimens
ROS/reactive nitrogen species (RNS) assay kits (OxiSelect; Cell Biolabs, San Diego, CA, USA) were utilised to quantify ROS levels in liver tissue, according to the manufacturer’s instructions. In brief, 50 μg fresh liver specimens in 1 ml PBS were homogenised (Precellys 24 homogeniser; Bertin Technologies, Versailles, France). Upon centrifugation, 50 μl tissue lysate was mixed with 50 μl catalyst solution and 100 μl DCFH solution (0.5 μl DCFH-DiOxyQ + 2 μl priming reagent + 97.5 μl stabilisation solution) for 30 min. The levels of 2′,7′-dichlorodihydrofluorescein (DCF) were quantified using fluorescence spectrometry with excitation/emission (Ex/Em) at 480/530 nm (FLUOstar OPTIMA; BMG Labtech, Ortenberg, Germany).
HSP60 cDNA transfection and RNA interference
pUSE-amp vectors coding full-length HSP60 (a gift from F.-S. Wang, Kaohsiung Chang Gung Memorial Hospital, Kaohsiung, Taiwan) were transfected into 2 × 105 HepG2 cells/well (in 6-well plates) for 48 h using Lipofectamine 3000 reagent (catalogue no. 3000015; Thermo Fisher Scientific, Waltham, MA, USA). In some experiments, HSP60 siRNA (catalogue no. sc-29351; Santa Cruz Biotechnology) and control siRNA (catalogue no. sc-37007; Santa Cruz Biotechnology) were transfected into cells for 4 h and then cells were incubated in basal medium for 48 h.
HSP60 immunofluorescence assay
Upon incubation with PA (200 μmol/l; catalogue no. 29558; Cayman Chemical), cells were fixed by paraformaldehyde and incubated in PBS with 0.1% (vol./vol.) Triton X-100 and 2% (vol./vol.) BSA. HSP60 in cells was stained using a primary HSP60 antibody and an Alexa Fluor-conjugated secondary antibody (Invitrogen; Carlsbad, CA, USA). Upon rinsing with PBS, intracellular lipid droplets were stained using 2 μg/ml Nile red (catalogue no. 72485; Merck KGaA, Darmstadt, Germany) and counterstained with 1 μg/ml DAPI (catalogue no. D5942; Merck KGaA). Immunostained cells were evaluated using fluorescence microscopy (LSM510; Carl Zeiss, Thornwood, NY, USA). Fluorescence intensity was measured using the ZEISS ZEN Blue software (Carl Zeiss).
Measurement of mitochondrial complex activity
Mitochondrial complex I and II activities in liver tissue were measured using the Abcam Complex I Enzyme Activity Assay (catalogue no. ab109721) and Complex II Enzyme Activity Assay (catalogue no. ab109908) and quantified, as previously described [22]. See ESM Methods for details.
Western blot analysis
After being transfected with pUSE-amp vector carrying HSP60 cDNA for 24 h, cells were treated with or without 15 nmol/l 3TYP (catalogue no. B2322; BioVision, Waltham, MA, USA) and incubated in 0.08 ml PBS with 0.8 mmol/l BSA (catalogue no. A7030; Merck KGaA; vehicle) or PA for another 24 h before total protein extraction. Cell lysates were extracted using Protein Extraction reagent (catalogue no. 78510; Thermo Fisher Scientific). Electrophoresis and immunoblotting of designated proteins, including HSP60, acyl-CoA dehydrogenase, long chain (ACADL), acyl-CoA cholesterin acyltransferase (ACAT1), acetyl coenzyme A acyltransferase 2 (ACAA2), carnitine palmitoyltransferase (CPT)1α, CPT2, SIRT3, AMP-activated protein kinase α (AMPKα), p-AMPKα, peroxisome proliferator-activated receptor, gamma, coactivator 1, α (PGC1α) and peroxisome proliferator-activated receptor α (PPARα) were conducted, as previously described [22]. The antibodies were used as per manufacturer’s instructions. Briefly, the membrane was blocked with 5% (vol./vol.) milk in TBS supplemented with 0.15% (vol./vol.) Tween-20 for 30 min. Membranes were then incubated with the indicated primary antibodies (1:1000 or 1:500 dilutions in TBS supplemented with 0.15% [vol./vol.] Tween-20 and 5% [vol./vol.] BSA) for 1 h, followed by secondary antibodies (Jackson ImmunoResearch, West Grove, PA, USA) conjugated with horseradish peroxidase (HRP; 1:5000 dilution in TBS supplemented with 0.15% [vol./vol.] Tween-20 and 5% [vol./vol.] milk) for 1 h. The relative intensity of designated protein bands was quantified using Image J software and normalised to the intensity of housekeeping β-actin protein (see Table 1).
Real-time PCR
Total RNA was isolated using the Quick-RNA Miniprep Kit (Zymo Research, Irvine, CA, USA) and reverse transcription was conducted using the GScript First-Strand Synthesis Kit (GeneDireX, Las Vegas, NV, USA), according to the manufacturers’ instructions. Upon mixing cDNA with primers and Fast SYBR Green Master Mix (Applied Biosystems, Foster City, CA, USA), PCR was conducted and Ct values were computed using the 7500 Fast Dx Real-Time PCR system (Applied Biosystems). Fold change of mRNA expression was calculated using \( {2}^{-\Delta {\mathrm{C}}_{\mathrm{t}}} \) and normalised to the housekeeping gene. Primer sequences for HSP60 and lipid metabolism-associated genes are listed in the Table 2 (relative mRNA expression was normalised to the housekeeping gene β-actin for these analyses), while primer sequences for the analysis of genes related to the unfolded protein response are described in ESM Methods (in which relative mRNA expression was normalised to the housekeeping gene Gapdh).
Statistical analyses
The data were analysed using GraphPad Prism 8.0 software (GraphPad Software, San Diego, CA, USA) and expressed as mean ± SD. Unpaired t tests were used to compare two groups. Two-way ANOVA and post hoc multiple comparison using the Tukey test were used to compare groups of WT and Hsp60-Tg mice fed with CD or HFD, as well as groups of HepG2 cells transfected with HSP60 cDNA or HSP60 siRNA and incubated in vehicle or PA. A p value <0.05 was considered statistically significant.
Results
Chaperone HSP60 levels are reduced in human fatty liver and murine DIO
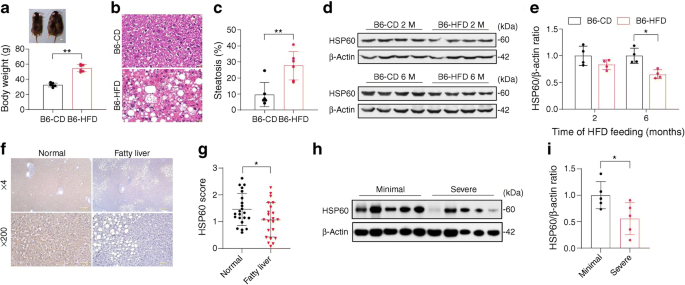
We first investigated whether HSP60 levels were changed in mice with DIO. The mice showed an obese body structure and gained body weight of approximately 20 g following HFD feeding for 6 months, as compared with CD-fed animals (Fig. 1a). Lipid accumulation in the hepatic tissue of HFD-fed mice was observed (Fig. 1b), together with a >20% increase in hepatic steatosis (Fig. 1c). Compared with CD-fed mice, decreased HSP60 levels were noted in hepatic tissue upon HFD consumption for 6 months, however this was not observed in mice fed an HFD for 2 months (Fig. 1d,e).
Analysis of hepatic steatosis and HSP60 expression in murine DIO and human fatty liver biopsies. (a) Overweight mice following CD (B6-CD) or HFD (B6-HFD) feeding for 6 months; scale bar, 1 cm. Data are mean ± SD (n=6). (b, c) Histopathological analysis showing liver adiposis, (b) together with severe hepatic steatosis in HFD-fed mice (c); scale bar, 20 μm. Data are presented as mean ± SD. (d, e) HSP60 levels in mouse liver following HFD consumption for 2 months (B6-HFD 2 M), HFD consumption for 6 months (B6-HFD 6 M), CD consumption for 2 months (B6-CD 2 M) or CD consumption for 6 months (B6-CD 6 M). In (e), relative HSP60 levels normalised to actin levels are presented as mean ± SD (n=4). (f) Weak HSP60 immunostaining in human fatty liver tissues; scale bar, 100 μm (high [×200] magnification) and 20 μm (low [×4] magnification). (g) Quantitative analysis of HSP60 distribution in fatty liver (n=22), as compared with normal liver (n=22). Data are presented as mean ± SD. (h, i) Lower HSP60 levels in severe fatty liver tissue (area of fat deposition >60%) as compared with those in minimal fatty liver (with area of fat deposition <5%). In (i), relative HSP60 levels were normalised to actin levels and are presented as mean ± SD (n=5). *p<0.05, **p<0.01
To investigate whether HSP60 loss could be recapitulated in human fatty liver, we conducted studies on 22 normal liver specimens and 22 fatty liver specimens from humans. In fatty liver, HSP60 immunoreaction, as evidenced by HSP60 scores, was less than in normal liver (Fig. 1f,g). Consistent with these findings, severe fatty livers (with >60% steatosis) had lower HSP60 levels than minimal fatty livers (<5% steatosis) (Fig. 1h,i), indicating that HSP60 loss is correlated with metabolic disorders.
Transgenic overexpression of HSP60 ameliorated DIO and increased dietary fat utilisation
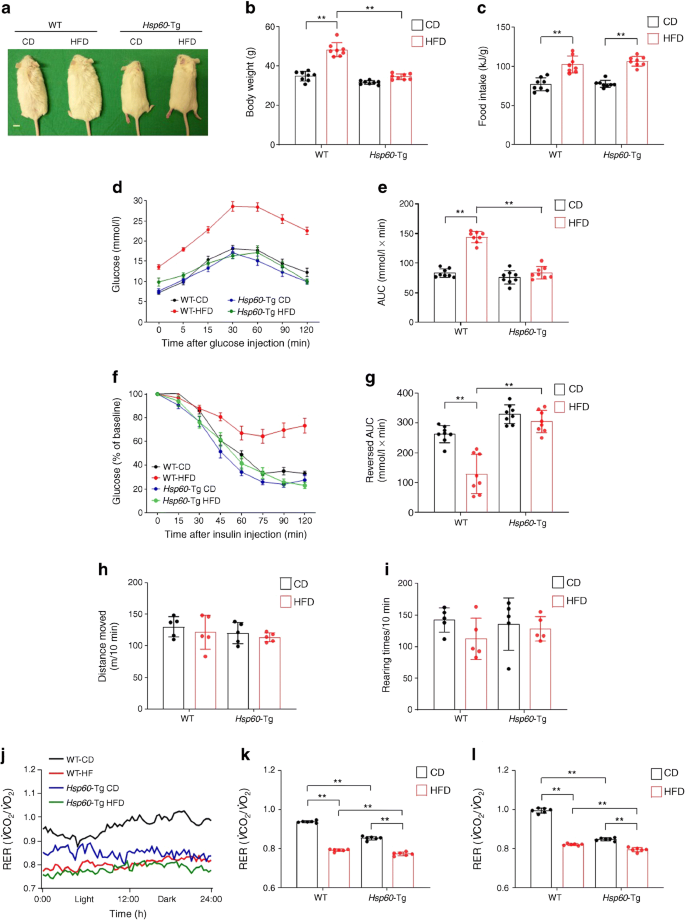
The observation that HSP60 was decreased in human fatty liver specimens and in mice with DIO prompted us to hypothesise that HSP60 overexpression could play a role in HFD-induced metabolic disorders. To investigate this, Hsp60-Tg or WT mice were fed with an HFD or CD for 6 months. The body size of the Hsp60-Tg mice remained similar upon HFD or CD feeding, whereas WT animals developed obesity upon HFD consumption (Fig. 2a). In addition, body weight of Hsp60-Tg mice was less than that of WT mice upon HFD feeding (Fig. 2b), despite food intake being similar between the two genotypes (Fig. 2c). Hsp60-Tg mice had higher glucose tolerance (Fig. 2d,e) and higher insulin sensitivity (Fig. 2f,g) than WT mice under HFD conditions, as evidenced by IPGTT and IPITT analyses, respectively.
Hsp60-Tg mice developed few signs of metabolic disorders upon HFD feeding. (a, b) HFD-fed WT mice showed an obese build (a) together with an increase in body weight (b), as compared with CD-fed mice, while HFD- and CD-fed Hsp60-Tg mice were similar; scale bar, 1 cm. Data are shown as mean ± SD (n=8). (c) Food intake was similar between WT and Hsp60-Tg mice upon CD or HFD feeding. Food intake measured over 2 days and normalised to body weight (kg); data are presented as mean ± SD (n=8). (d) Analysis of glucose tolerance by IPGTT in WT and Hsp60-Tg mice fed with CD or HFD. (e) The AUC of serum glucose levels at 120 min post IPGTT was reduced in HFD-fed Hsp60-Tg mice vs HFD-fed WT mice. Data are presented as mean ± SD. (f) Analysis of insulin tolerance profiles by IPITT in WT and Hsp60-Tg mice fed with CD or HFD. (g) The reversed AUC of serum glucose levels at 120 min post IPTTT demonstrates that HFD-induced insulin resistance was improved in Hsp60-Tg mice vs WT mice. Data are presented as mean ± SD (n=8). (h, i) Physical activity, as measured by moving distance (h) and rearing time (i), was similar in WT and Hsp60-Tg mice upon CD or HFD feeding. Data are presented as mean ± SD (n=5). (j) RER profiles of WT and Hsp60-Tg mice fed a CD or HFD. (k, l) Hsp60-Tg mice had a lower RER than WT mice upon living in light (k) and dark (l) conditions. Data are presented as mean ± SD (n=6). **p<0.01
Physical activity including moving distance and rearing activity, was similar between Hsp60-Tg and WT mice (Fig. 2h,i). The WT mice had a lower RER profile upon HFD feeding vs CD feeding. Of interest, the profiles of the Hsp60-Tg mice was low with both HFD and CD consumption (Fig. 2j). In addition, the RER profiles of Hsp60-Tg mice were lower than those of WT mice when accommodated in light (Fig. 2k) and dark (Fig. 2l) conditions, suggesting that Hsp60-Tg mice may utilise fatty acids over carbohydrates.
HSP60 repressed HFD-induced adiposis, adipokines and inflammation
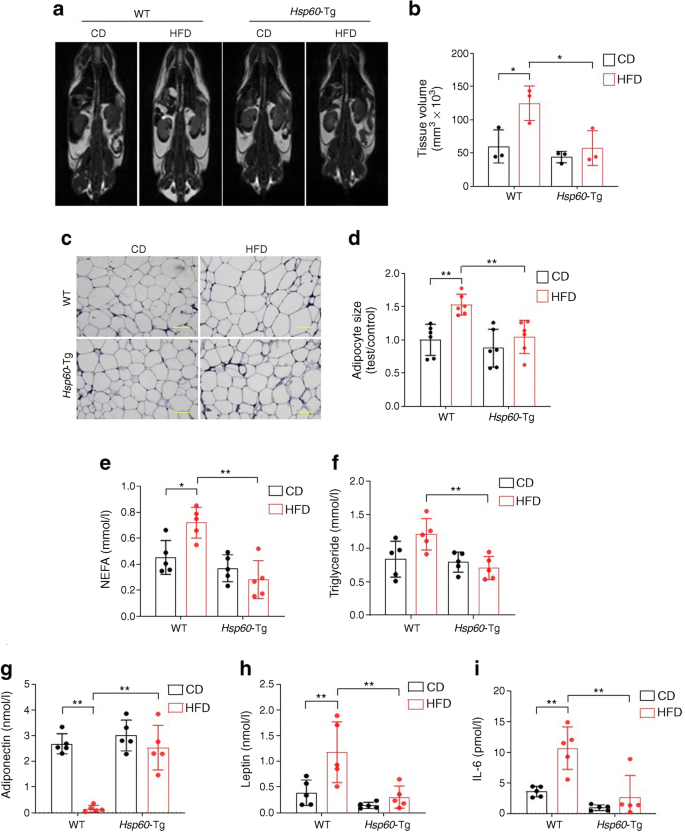
We evaluated whether HSP60 overexpression altered fat metabolism. MRI images revealed that body fat overdevelopment (Fig. 3a) and fat tissue volume (Fig. 3b) were lower in Hsp60-Tg mice vs WT mice following HFD consumption. An abundance of enlarged adipocytes (Fig. 3c), together with increased adipocyte size (Fig. 3d), high serum NEFA levels (Fig. 3e) and increased serum triglyceride levels (Fig. 3f; not significant) were observed in the HFD-fed vs CD-fed WT mice; however, these effects were limited in HFD-fed Hsp60-Tg mice. Lower serum adiponectin levels (Fig. 3g) and higher serum leptin (Fig. 3h) and IL-6 (Fig. 3i) levels were observed in HFD-fed WT mice, as compared with CD-fed WT mice. However, HFD-mediated serum leptin and IL-6 overproduction were obliterated in the Hsp60-Tg mice. In addition, the changes in adiponectin levels observed in WT mice upon HFD feeding vs CD feeding were not visible in Hsp60-Tg mice, suggesting that HSP60 overexpression improved HFD-mediated adiposis and adipokine dysregulation.
HSP60 overexpression reduced HFD-mediated body adiposis and adipokine dysregulation. (a) MRI analysis of body fat distribution. (b) HFD-induced increases in total fat volume were reduced in Hsp60-Tg mice vs WT mice (n=3). (c) Histological analysis of visceral fat in WT and Hsp60-Tg mice fed with a CD or HFD; scale bar, 100 μm. (d) HFD-mediated increases in adipocyte size were compromised in Hsp60-Tg mice (n=6). (e, f) HFD-induced increases in serum NEFA (e) and triglyceride (f) levels were reversed in Hsp60-Tg mice (n=5). (g–i) HFD-induced reductions in serum adiponectin levels (g) and increases in serum leptin (h) and IL-6 (i) levels were abolished in Hsp60-Tg mice. Data are presented as mean ± SD (n=5 mice). *p<0.05, **p<0.01
HSP60 alleviated HFD-mediated hepatic steatosis, oxidative stress and M1/M2 macrophage switch
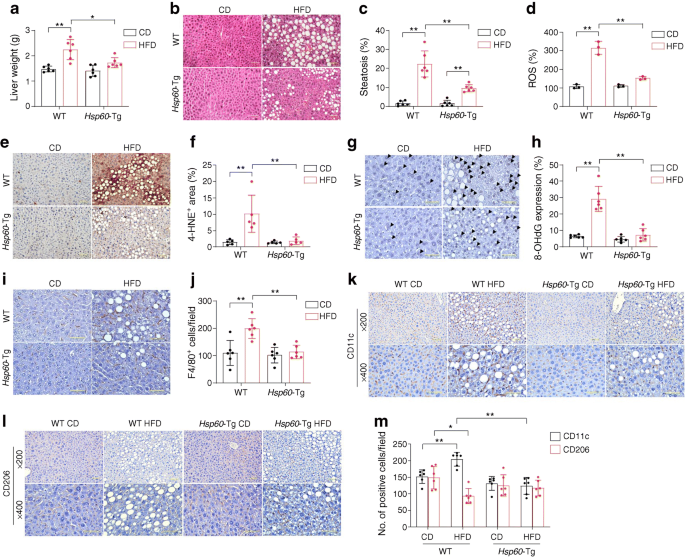
Increased liver weight was observed in the HFD-fed WT mice (Fig. 4a) but not in HFD-fed Hsp60-Tg mice vs CD-fed counterparts. Compared with CD-fed mice of the same genotype, HFD-fed WT mice developed hepatic steatosis, as determined by histopathology, together with a 20% increase in steatosis area (Fig. 4b,c). In contrast, HFD-mediated liver steatosis was ameliorated in the Hsp60-Tg mice. Furthermore, HFD upregulated ROS production (Fig. 4d), and enhanced 4-HNE (Fig. 4e,f) and 8-OHdG accumulation (Fig. 4g,h) in the liver tissue of the WT mice, as compared with CD-fed controls. In contrast, however, the hepatic tissue of HFD-fed Hsp60-Tg mice was shown to have lower levels of these markers of lipid oxidation and DNA oxidative damage.
HSP60 overexpression attenuated HFD-mediated liver steatosis, oxidative damage and M1/M2 macrophage dysregulation. (a) HFD-induced increases in liver weight were repressed in Hsp60-Tg mice (n=6). Data are presented as mean ± SD. (b, c) Histological images (b) and quantification (c) showing reduced liver steatosis in HFD-fed Hsp60-Tg mice than HFD-fed WT mice; scale bar, 100 μm. Data are presented as mean ± SD. (d) HFD-induced ROS overproduction was improved in Hsp60-Tg mice (n=3 mice). Data are presented as mean ± SD. (e) 4-HNE immunostaining in liver tissues following CD or HFD feeding; scale bar, 100 μm. (f) 4-HNE-immunostained area in HFD-fed Hsp60-Tg mice was reduced when compared with HFD-fed WT mice (n=5). Data are presented as mean ± SD. (g) 8-OHdG immunoreaction (arrowheads) in liver tissues; scale bar, 20 μm. (h) Hsp60-Tg mice had less 8-OHdG-immunostained cells as compared with WT mice following HFD feeding (n=6). Data are presented as mean ± SD. (i) F4/80-immunostained macrophages in liver tissues; scale bar, 20 μm. (j) HSP60 overexpression attenuated the HFD-induced increase in F4/80-immunostained cells. Data are presented as mean ± SD (n=6). (k, l) CD11c-immunostained M1 macrophages (k) and CD206-immunostained M2 macrophages (l) in liver tissue; scale bar, 100 μm (higher magnification) and 20 μm (lower magnification). (m) The HFD-induced increase in M1 macrophages was repressed in Hsp60-Tg mice. Data are presented as mean ± SD (n=6). *p<0.05, **p<0.01
A higher F4/80-positive macrophage count was detected in liver tissue from WT mice fed with an HFD as compared with liver tissues from HFD-fed Hsp60-Tg mice, indicating that HSP60 overexpression reduces HFD-induced macrophage infiltration (Fig. 4i,j). To further clarify macrophage phenotype, we utilised M1 macrophage marker (CD11c) and M2 macrophage marker (CD206) antibodies to investigate macrophage phenotype profiles in liver tissues. A higher CD11c-positive proinflammatory macrophage count and lower CD206-positive anti-inflammatory macrophage count were observed in liver tissues of WT mice fed with an HFD as compared with CD-fed counterparts, whereas HFD-fed Hsp60-Tg had similar levels of CD11c and CD206 as CD-fed Hsp60-Tg mice (Fig. 4k–m). HFD-mediated macrophage infiltration and dysregulation of M1/M2 macrophage polarisation were ameliorated in the Hsp60-Tg mice. HSP60-associated repression of macrophage infiltration was also observed in adipose tissue, as demonstrated by the presence of a plethora of CD11c-immunostained macrophages adjacent to crown-like structures (CLS), and reduced CLS-adjacent CD206-immunostained macrophages in adipose samples from HFD-fed WT mice as compared with CD-fed controls (ESM Fig. 1a,b). The effects of HFD feeding on the number of CLS-adjacent CD206-positive cells were reversed in HFD-fed Hsp60-Tg mice vs CD-fed controls (ESM Fig. 1a,b), suggesting that HSP60 provides protection against metabolic inflammation.
HSP60 helped to maintain the mitochondrial unfolded protein response and improve electron transport chain function
Dysregulation of the mitochondrial unfolded protein response (mtUPR) is a common feature in the development of metabolic disorders [23]. We thus examined whether HSP60 affected the mtUPR in HFD-induced fatty livers or adipose tissue. HFD feeding increased the expression of key mtUPR markers, including activating transcription factor (ATF)4, ATF5, C/EBP homologous protein (CHOP) and LON peptidase 1 (LONP1) in liver tissue (ESM Fig. 2a) and visceral fat (ESM Fig. 2b) from WT mice, as compared with CD-fed WT mice. With the exception of Lonp1 in visceral fat, the expression of these genes in HFD-fed Hsp60-Tg mice was lower than in HFD-fed WT mice. In addition, in WT mice, HFD feeding promoted mitochondrial complex I (ESM Fig. 2c) and complex II (ESM Fig. 2d) activity in the liver but reduced this in visceral fat, as compared with CD-fed WT mice. Of interest, the activity of mitochondrial complex I and II in Hsp60-Tg mice was higher than in WT mice upon CD or HFD feeding, although the increase in complex II activity in visceral fat from CD-fed Hsp60-Tg vs WT mice was not significant. These data indicate that HSP60 overexpression may reduce excess nutrient-induced mitochondrial stress and enhanced mitochondrial electron transport chain (ETC) activity.
Forced HSP60 expression promoted fatty acid oxidation in HepG2 cells and Hsp60-Tg mice
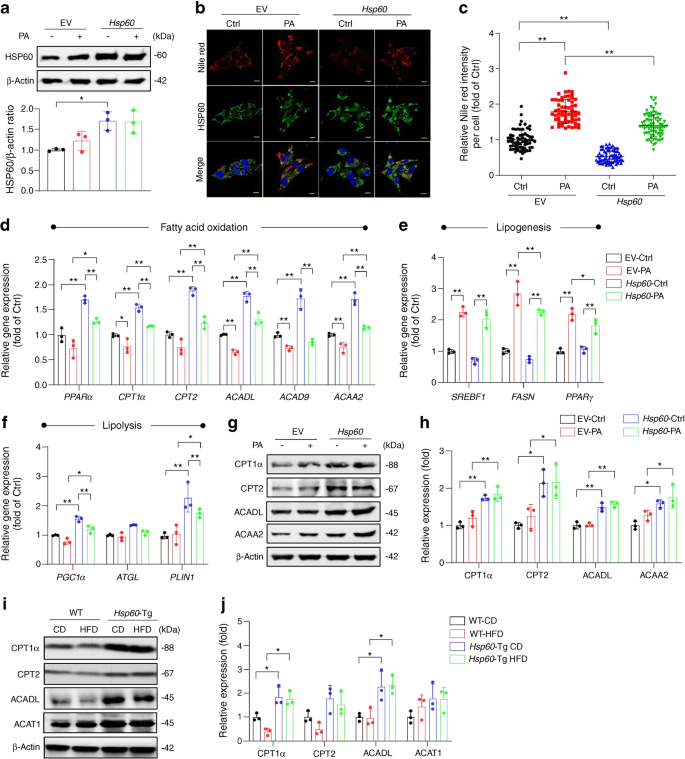
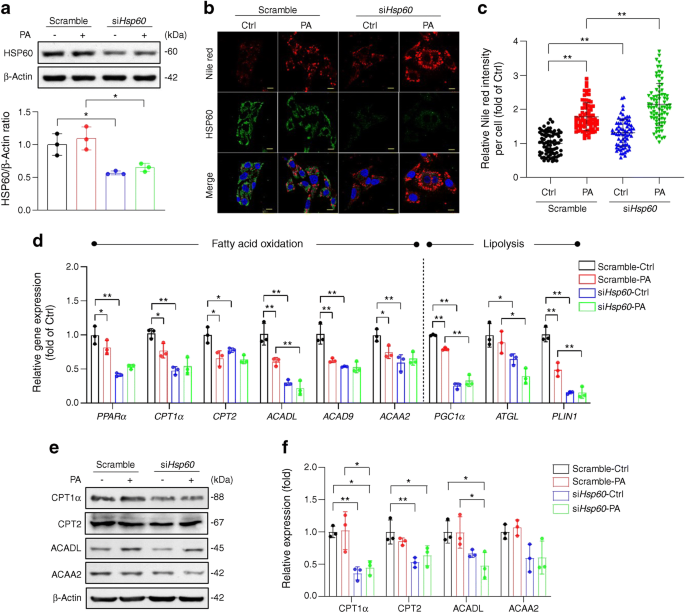
To investigate how HSP60 inhibited hepatic steatosis, we incubated HepG2 cells in PA as an in vitro HFD-induced fatty liver model [24], and increased HSP60 expression in cells by transfecting them with full-length HSP60 cDNA (Fig. 5a). As indicated by Nile red staining, there was an abundance of intracellular lipid droplets, along with weak HSP60 immunofluorescence in empty vector (EV)-transfected cells upon PA incubation (Fig. 5b). Forced HSP60 expression attenuated PA-induced lipid formation (Fig. 5b,c). In EV-transfected cells, PA repressed expression of fatty acid beta oxidation markers, including PPARα, CPT1α, CPT2, ACADL, ACAD9 and ACAA2 (Fig. 5d; changes in PPARα and CPT2 were non-significant) and enhanced the expression of lipogenesis markers SREBF1, FASN and PPARγ (Fig. 5e), whereas the expression of lipolysis markers PGC1α, ATGL and PLIN1 were unaffected (Fig. 5f). Of note, compared with EV-vector transfected cells, forced HSP60 expression promoted expressions of fatty acid beta oxidation markers and the lipolysis markers PGC1α and PLIN1, while lipogenesis markers were unaffected. HSP60 overexpression also attenuated PA-induced dysregulation of several genes associated with fatty acid oxidation, lipolysis and lipogenesis, as compared with EV-transfected cells incubated with PA (Fig. 5d–f). Consistent with mRNA expression, protein levels of CPT1α, CPT2, ACADL and ACAA2 were elevated in HSP60-transfected HepG2 cells compared with EV-transfected controls (Fig. 5g,h), of which CPT1α and ACADL were significantly increased in liver tissues of Hsp60-Tg mice vs WT mice upon CD or HFD feeding (Fig. 5i,j).
Forced HSP60 expression reduced lipid accumulation and promoted fatty acid oxidation. (a) Increased HSP60 levels in HepG2 cells upon HSP60 transfection. Data are presented as mean ± SD. (b) Fluorescence images showing Nile red-stained oil droplets and HSP60 immunofluorescence in HepG2 cells; scale bar, 10 μm. (c) Forced HSP60 expression attenuated PA-induced, Nile red-stained oil production. Data are presented as mean ± SD (n=60–70 cells). (d–f) Expression of key markers of fatty acid oxidation (d), lipogenesis (e) and lipolysis (f) in HepG2 cells, quantified using RT-PCR. Key in (e) also applies to (d) and (f). Data are presented as mean ± SD (n=3 experiments). (g, h) Effects of forced HSP60 expression and PA incubation on CPT1α, CPT2, ACADL and ACAA2 levels in HepG2 cells. (i, j) Changes in CPT1α, CPT2, ACADL and acyl-CoA cholesterin acyltransferase (ACAT1) levels in liver tissue from WT and Hsp60-Tg mice following CD and HFD feeding mice. Data are presented as mean ± SD. Ctrl, control *p<0.05, **p<0.01
HSP60 loss enhanced PA-induced lipid formation and decreased fatty acid oxidation
HSP60 knockdown by HSP60 siRNA in HepG2 cells (Fig. 6a) upregulated Nile red-stained oil droplet formation, which was further exaggerated upon PA incubation (Fig. 6b,c). HSP60 interference reduced basal expression of fatty acid oxidation markers and lipolysis markers (Fig. 6d). Compared with basal levels, PA incubation also reduced expression of fatty acid oxidation markers and lipolysis markers, except of ATGL (Fig. 6d). Compared with cells treated with scramble siRNA and PA, HSP60 interference with PA treatment significantly reduced expression of the fatty acid oxidation marker ACADL and the lipolysis markers PGC1ɑ, ATGL and PLIN1 (Fig. 6d). Downregulation of protein levels of CPT1α, CPT2 and ACADL in siHSP60-transfected cells vs scramble-transfected controls was evident, although ACAA2 was not significantly different across any of the treatment groups (Fig. 6e,f). These results indicate that HSP60 inhibits lipid overproduction by regulating fatty acid oxidation and lipolysis.
HSP60 knockdown aggravated lipid accumulation and decreased fatty acid oxidation in PA-treated HepG2 cells. (a–c) Hsp60 interference repressed HSP60 levels (a) and HSP60 immunofluorescence (b; scale bar, 10 μm) and enhanced (Nile red-stained) oil droplet production (c). Data are presented as mean ± SD (n=60–70). (d) Effects of HSP60 knockdown and PA incubation on expression of key markers of fatty acid oxidation and lipolysis, quantified using RT-PCR. Data are presented as mean ± SD (n=3 experiments). (e, f) CPT1α, CPT2, ACADL and ACAA2 levels in HepG2 cells upon HSP60 knockdown and/or PA incubation. Data are presented as mean ± SD (n=3 experiments). Ctrl, control; *p<0.05, **p<0.01
SIRT3 signalling mediated HSP60-associated promotion of fatty acid oxidation in Hsp60-Tg mice and HepG2 cells
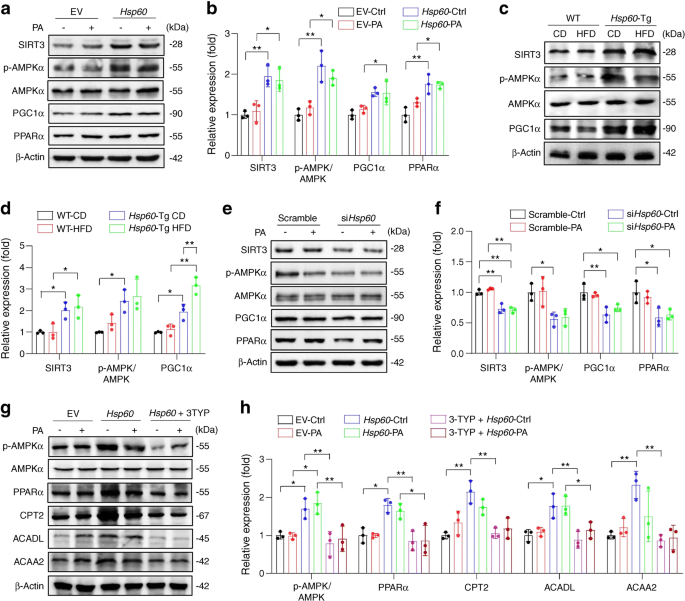
We aimed to identify the molecular mechanism underlying HSP60-associated repression of liver adiposity. SIRT3 is known to regulate AMPK/ PGC1α/ PPARα signalling and control lipid metabolism [25]. Forced HSP60 expression increased SIRT3, PPARα and p-AMPK levels, with an upward trend (p>0.05) in PGC1α expression in HepG2 cells, while levels of these proteins were unaltered in EV-transfected cells upon PA incubation vs EV-transfected controls (Fig. 7a,b). The increase in levels of these proteins also occurred in liver tissue from Hsp60-Tg mice upon CD or HFD feeding vs WT counterparts, although the increase in p-AMPK was not significantly different between HFD-fed WT and Hsp60-Tg mice (Fig. 7c,d). In HepG2 cells, HSP60 knockdown caused reductions in SIRT3, PGC1α, PPARα and p-AMPK levels (Fig. 7e,f). Furthermore, pharmacological inhibition of SIRT3 using the inhibitor 3-(1H-1,2,3-triazol-4-yl) pyridine (3TYP) inhibited HSP60 overexpression-mediated promotion of p-AMPK and PPARα, while also decreasing levels of fatty acid oxidation markers (Fig. 7g,h). Our results suggest that SIRT3 mediated the HSP60-associated protection of fatty acid oxidation observed in PA-stressed hepatic cells.
SIRT3/AMPK/PGC1α signalling mediated HSP60-associated promotion of fatty acid oxidation. (a, b) Effects of forced HSP60 expression and PA incubation on SIRT3, p-AMPKα, PGC1α and PPARα levels. (c, d) Changes in SIRT3, p-AMPKα and PGC1α levels in liver tissue of WT and Hsp60-Tg mice upon CD or HFD consumption. (e, f) HSP60 knockdown downregulated SIRT3, p-AMPK, PGC-1α and PPARα levels in HepG2 cells. (g, h) SIRT3 inhibition by 3TYP inhibited HSP60-induced enhancement of protein levels of p-AMPKα, PPARα and ACADL in HepG2 cells incubated with PA. Data are presented as mean ± SD (n=3 experiments). Ctrl, control. *p<0.05, **p<0.01
Discussion
Overnutrition aggravates a variety of unfavourable outcomes, including increased adiposity [26], lipotoxicity [27], chronic inflammation [28] and oxidative stress [29], which accelerate hepatocyte dysfunction in the development of NAFLD, a prominent feature of metabolic disorders [30]. To maintain energy metabolism in the hepatic microenvironment in order to inhibit NAFLD is highly demanding. HSPs are highly conserved molecules that modulate intercellular homeostasis, adapting external stresses to maintain cellular activity [31]. HSP60 is indispensable for protecting against protein aggregation or misfolding in the hepatic microenvironment [32]. Our study demonstrates the profound protective effects of mitochondrial HSP60, which acts to inhibit the development of DIO and fatty liver by promoting lipid catabolism. In addition, we provide valuable insight into the function of SIRT3, identifying it as a functional effector of HSP60.
Our investigation reveals that HSP60 loss is associated with human hepatic steatosis and HFD-induced fatty liver in mice. Our results are consistent with previous studies showing that HSP60 mRNA and protein expression are decreased in obese humans with diabetes [33], while low HSP60 protein levels are found in individuals with type 2 diabetes [34]. To clarify the role that this mitochondrial chaperonin may play in HFD-mediated metabolic dysregulation, we studied Hsp60-Tg mice. Eight-week-old Hsp60-Tg FVB mice fed with an HFD for 6 months had lower levels of indicators of metabolic disorders (which include overweight status, glucose intolerance, insulin resistance, hypertriglyceridaemia, increased adiposity and hepatic steatosis). Previous studies have shown that 4-week-old C57BL/6N mice deficient in ubiquitous HSP60 had less obesity and fat mass than control mice upon HFD feeding for 16 weeks [35], whereas mice with hypothalamus-specific HSP60 knockout were insulin resistant [8]. We, thus, speculated that the role of HSP60 in metabolic disease development may be tissue context-dependent, and that mice with different genetic backgrounds or ages may have differing energy demands or fat metabolism capacities.
In our study, RER was low in both CD- or HFD-fed Hsp60-Tg mice. The finding that low RER occurred simultaneously to low lipid accumulation in adipose and hepatic tissues indicates that HSP60 may facilitate the utilisation of dietary fat during energy expenditure. Impaired fatty acid oxidation is a hallmark of obesity development [36], while promotion of lipolysis prevents increased adiposity and body weight gain [37, 38]. Here, Hsp60-Tg mice exhibited high fatty acid oxidation in the liver. The in vitro models in this study further revealed that upregulation of HSP60 compromised PA-induced loss of fatty acid oxidation and lipolysis capacity in HepG2 cells, while HSP60 loss worsened PA-induced inhibition of lipid catabolism. Collectively, our findings provide valuable insight into the role of HSP60 in inhibiting body and hepatic fat overburden following overconsumption of dietary fat.
Our investigation reveals that HFD-induced aggravation of the unfolded protein response is reversed in Hsp60-Tg mice. Unfolded protein response dysregulation in the endoplasmic reticulum enhances intracellular stress, accelerating fat accumulation in HFD-fed mice [39]. Moreover, the mtUPR is critical in adapting to overnutrition-induced stress, improving hepatic fatty acid oxidation and mitochondrial complex activities [40]. Meanwhile, mitochondrial HSP60 is a key component of the mtUPR, improving metabolic stress and cell survival [41]. This study demonstrates the mito-protective role of HSP60 in facilitating liver and adipose tissue to accommodate HFD-induced metabolic stress. In Hsp60-Tg mice, following chronic excess dietary fat intake, few signs of tissue deterioration (which include lipotoxicity, and lipid and DNA oxidation) were observed, along with reduced macrophage infiltration and M1/M2 macrophage dysregulation, and lower inflammatory cytokine secretion, further confirming the critical function that HSP60 plays in protecting against hepatic tissue dysfunction.
Key regulators of mitochondrial biogenesis, including SIRT3, AMPK and PGC1α signalling molecules, remained high in HFD-fed Hsp60-Tg mice vs WT counterparts. SIRT3 is an NAD+-dependent mitochondrial protein deacetylase, which regulates energy metabolism and oxidative stress [14, 42,43,44], and ameliorates fatty liver, lipogenesis and lipotoxicity [15, 16, 45]. In addition, SIRT3 is a mitochondrial fidelity protein that senses nutrients and redox status. SIRT3 overexpression promotes the breakdown of lipid droplets via chaperon-mediated autophagy [16]. It also enhances fatty acid β-oxidation [14, 16, 46] and represses lipogenesis [16] and lipotoxicity [45], in addition to working with AMPK/PGC-1α signalling to mitigate oxidative stress and cell death [47,48,49]. In our HepG2 cell model, HSP60 overexpression promoted SITR3 signalling, and elevated AMPK, PGC1α and PPARα levels. The promotion of fatty acid oxidation and lipolysis following HSP60 overexpression were repressed upon introduction of the SIRT3 inhibitor 3TYP, suggesting that SIRT3 mediates the HSP60-associated actions on fatty acid metabolism. On a molecular level, our study provides valuable insight into how HSP60 impedes the development of DIO and hepatic steatosis.
Mounting evidence has revealed that SIRT3 modulates mitochondrial chaperone complex activity through deacetylating HSPs [14, 50], while HSP60 loss results in misfolding of SIRT3 leading to it being degraded more quickly. As previously reported, mice with cardiac-specific Hsp60 knockout exhibited low mitochondrial bioenergetics and SIRT3 signalling, together with high ROS levels, leading to the development of dilated cardiopathy [51]. SIRT3 may interplay with HSP60 to orchestrate mitochondrial metabolism, thereby promoting fat utilisation.
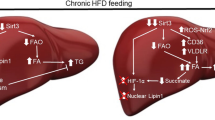
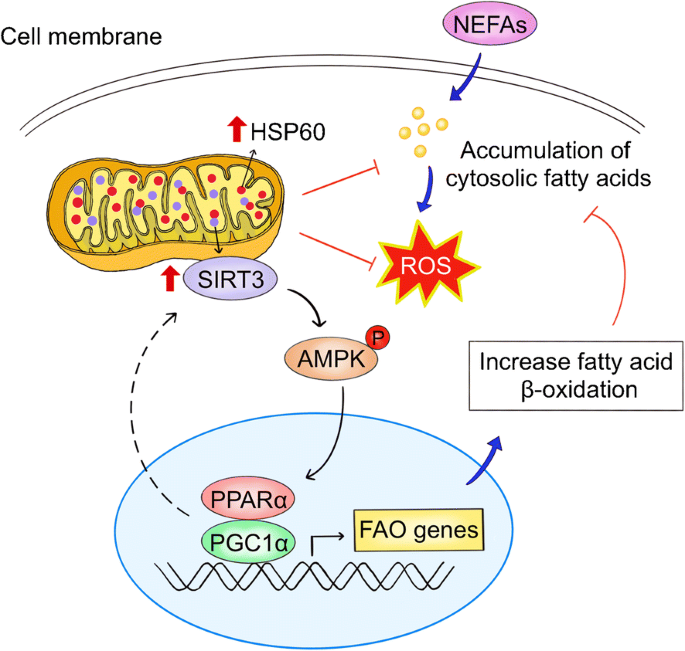
Our investigations collectively demonstrate that HSP60 promotes fatty acid oxidation and lipolysis, while inhibiting ROS, lipotoxicity and mitochondrial dysfunction, thereby preventing increased adiposity and fatty liver development. SIRT3 orchestrates AMPK and PGC-1α pathways, contributing to the HSP60-associated attenuation of HFD-mediated obesity and hepatic steatosis (Fig. 8). The findings herein indicate that manipulation of mitochondrial HSP60 may be a promising direction for the development of therapeutic interventions for NAFLD and type 2 diabetes.
Data availability
Data are available from the corresponding author upon reasonable request.
Abbreviations
- ACAA2:
-
Acetyl coenzyme A acyltransferase 2
- ACADL:
-
Acyl-CoA dehydrogenase, long chain
- AMPK:
-
AMP-activated protein kinase
- ATP:
-
Activating transcription factor
- CD:
-
Chow diet
- CPT:
-
Carnitine palmitoyltransferase
- DIO:
-
High-fat-diet-induced obesity
- EV:
-
Empty vector
- HFD:
-
High-fat diet
- 4-HNE:
-
4-Hydroxy-2-nonenal
- HSP:
-
Heat shock protein
- LONP1:
-
LON peptidase 1
- mtUPR:
-
Mitochondrial unfolded protein response
- NAFLD:
-
Non-alcoholic fatty liver disease
- 8-OHdG:
-
8-Hydroxy-2′-deoxyguanosine
- PA:
-
Palmitic acid
- PGC-1α:
-
Peroxisome proliferator-activated receptor, gamma, coactivator 1, α
- PPARα:
-
Peroxisome proliferator-activated receptor α
- RER:
-
Respiratory exchange ratio
- RNAi:
-
RNA interference
- ROS:
-
Reactive oxygen species
- SIRT3:
-
Sirtuin 3
- Tg:
-
Transgenic
- 3TYP:
-
3-(1H-1,2,3-triazol-4-yl) pyridine
- WT:
-
Wild-type
References
Targher G, Byrne CD, Tilg H (2020) NAFLD and increased risk of cardiovascular disease: clinical associations, pathophysiological mechanisms and pharmacological implications. Gut 69(9):1691–1705. https://doi.org/10.1136/gutjnl-2020-320622
Rolo AP, Teodoro JS, Palmeira CM (2012) Role of oxidative stress in the pathogenesis of nonalcoholic steatohepatitis. Free Radic Biol Med 52(1):59–69. https://doi.org/10.1016/j.freeradbiomed.2011.10.003
Begriche K, Igoudjil A, Pessayre D, Fromenty B (2006) Mitochondrial dysfunction in NASH: causes, consequences and possible means to prevent it. Mitochondrion 6(1):1–28. https://doi.org/10.1016/j.mito.2005.10.004
Meex RCR, Blaak EE (2021) Mitochondrial dysfunction is a key pathway that links saturated fat intake to the development and progression of NAFLD. Mol Nutr Food Res 65(1):e1900942. https://doi.org/10.1002/mnfr.201900942
Caruso Bavisotto C, Alberti G, Vitale AM et al (2020) Hsp60 post-translational modifications: functional and pathological consequences. Front Mol Biosci 7:95. https://doi.org/10.3389/fmolb.2020.00095
Juwono J, Martinus RD (2016) Does Hsp60 provide a link between mitochondrial stress and inflammation in diabetes mellitus? J Diabetes Res 2016:8017571. https://doi.org/10.1155/2016/8017571
Liyanagamage D, Martinus RD (2020) Role of mitochondrial stress protein HSP60 in diabetes-induced neuroinflammation. Mediators Inflamm 2020:8073516. https://doi.org/10.1155/2020/8073516
Kleinridders A, Lauritzen HP, Ussar S et al (2013) Leptin regulation of Hsp60 impacts hypothalamic insulin signaling. J Clin Invest 123(11):4667–4680. https://doi.org/10.1172/JCI67615
Docrat TF, Nagiah S, Naicker N, Baijnath S, Singh S, Chuturgoon AA (2020) The protective effect of metformin on mitochondrial dysfunction and endoplasmic reticulum stress in diabetic mice brain. Eur J Pharmacol 875:173059. https://doi.org/10.1016/j.ejphar.2020.173059
Wilson-Fritch L, Nicoloro S, Chouinard M et al (2004) Mitochondrial remodeling in adipose tissue associated with obesity and treatment with rosiglitazone. J Clin Invest 114(9):1281–1289. https://doi.org/10.1172/JCI21752
Elkhwanky MS, Hakkola J (2018) Extranuclear sirtuins and metabolic stress. Antioxid Redox Signal 28(8):662–676. https://doi.org/10.1089/ars.2017.7270
Rardin MJ, Newman JC, Held JM et al (2013) Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways. Proc Natl Acad Sci U S A 110(16):6601–6606. https://doi.org/10.1073/pnas.1302961110
Lombard DB, Alt FW, Cheng HL et al (2007) Mammalian Sir2 homolog SIRT3 regulates global mitochondrial lysine acetylation. Mol Cell Biol 27(24):8807–8814. https://doi.org/10.1128/MCB.01636-07
Lu Z, Chen Y, Aponte AM, Battaglia V, Gucek M, Sack MN (2015) Prolonged fasting identifies heat shock protein 10 as a Sirtuin 3 substrate: elucidating a new mechanism linking mitochondrial protein acetylation to fatty acid oxidation enzyme folding and function. J Biol Chem 290(4):2466–2476. https://doi.org/10.1074/jbc.M114.606228
Barroso E, Rodriguez-Rodriguez R, Zarei M et al (2020) SIRT3 deficiency exacerbates fatty liver by attenuating the HIF1alpha-LIPIN 1 pathway and increasing CD36 through Nrf2. Cell Commun Signal 18(1):147. https://doi.org/10.1186/s12964-020-00640-8
Zhang T, Liu J, Shen S, Tong Q, Ma X, Lin L (2020) SIRT3 promotes lipophagy and chaperon-mediated autophagy to protect hepatocytes against lipotoxicity. Cell Death Differ 27(1):329–344. https://doi.org/10.1038/s41418-019-0356-z
Fernandez-Marcos PJ, Jeninga EH, Canto C et al (2012) Muscle or liver-specific Sirt3 deficiency induces hyperacetylation of mitochondrial proteins without affecting global metabolic homeostasis. Sci Rep 2:425. https://doi.org/10.1038/srep00425
Ko JY, Sun YC, Li WC, Wang FS (2016) Chaperonin 60 regulation of SOX9 ubiquitination mitigates the development of knee osteoarthritis. J Mol Med 94(7):755–769. https://doi.org/10.1007/s00109-016-1422-3
National Research Council (US) Subcommittee on Laboratory Animal Nutrition. Nutrient Requirements of Laboratory Animals: Fourth Revised Edition, 1995. National Academies Press, Washington, DC
Wang PW, Kuo HM, Huang HT et al (2014) Biphasic response of mitochondrial biogenesis to oxidative stress in visceral fat of diet-induced obesity mice. Antioxid Redox Signal 20(16):2572–2588. https://doi.org/10.1089/ars.2013.5334
Takahashi Y, Fukusato T (2014) Histopathology of nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. World J Gastroenterol 20(42):15539–15548. https://doi.org/10.3748/wjg.v20.i42.15539
Kuo HM, Weng SW, Chang AY et al (2016) Altered mitochondrial dynamics and response to insulin in cybrid cells harboring a diabetes-susceptible mitochondrial DNA haplogroup. Free Radic Biol Med 96:116–129. https://doi.org/10.1016/j.freeradbiomed.2016.04.019
Theurey P, Rieusset J (2017) Mitochondria-associated membranes response to nutrient availability and role in metabolic diseases. Trends Endocrinol Metab 28(1):32–45. https://doi.org/10.1016/j.tem.2016.09.002
Krishnan S, Ding Y, Saedi N et al (2018) Gut microbiota-derived tryptophan metabolites modulate inflammatory response in hepatocytes and macrophages. Cell Rep 23(4):1099–1111. https://doi.org/10.1016/j.celrep.2018.03.109
Han Y, Zhou S, Coetzee S, Chen A (2019) SIRT4 and its roles in energy and redox metabolism in health, disease and during exercise. Front Physiol 10:1006. https://doi.org/10.3389/fphys.2019.01006
Azzu V, Vacca M, Virtue S, Allison M, Vidal-Puig A (2020) Adipose tissue-liver cross talk in the control of whole-body metabolism: implications in nonalcoholic fatty liver disease. Gastroenterology 158(7):1899–1912. https://doi.org/10.1053/j.gastro.2019.12.054
Tilg H, Adolph TE, Dudek M, Knolle P (2021) Non-alcoholic fatty liver disease: the interplay between metabolism, microbes and immunity. Nat Metab 3(12):1596–1607. https://doi.org/10.1038/s42255-021-00501-9
Okubo S, Shindoh J, Kobayashi Y, Matsumura M, Hashimoto M (2022) Adhesions as a risk factor for postoperative morbidity in patients undergoing repeat hepatectomy and the potential efficacy of adhesion barriers. J Hepatobiliary Pancreat Sci 29(6):618–628. https://doi.org/10.1002/jhbp.1047
Chen Z, Tian R, She Z, Cai J, Li H (2020) Role of oxidative stress in the pathogenesis of nonalcoholic fatty liver disease. Free Radic Biol Med 152:116–141. https://doi.org/10.1016/j.freeradbiomed.2020.02.025
Targher G, Lonardo A, Byrne CD (2018) Nonalcoholic fatty liver disease and chronic vascular complications of diabetes mellitus. Nat Rev Endocrinol 14(2):99–114. https://doi.org/10.1038/nrendo.2017.173
Dubrez L, Causse S, Borges Bonan N, Dumetier B, Garrido C (2020) Heat-shock proteins: chaperoning DNA repair. Oncogene 39(3):516–529. https://doi.org/10.1038/s41388-019-1016-y
Mattoo RU, Goloubinoff P (2014) Molecular chaperones are nanomachines that catalytically unfold misfolded and alternatively folded proteins. Cell Mol Life Sci 71(17):3311–3325. https://doi.org/10.1007/s00018-014-1627-y
Khadir A, Kavalakatt S, Cherian P et al (2018) Physical exercise enhanced heat shock protein 60 expression and attenuated inflammation in the adipose tissue of human diabetic obese. Front Endocrinol 9:16. https://doi.org/10.3389/fendo.2018.00016
Houzelle A, Jorgensen JA, Schaart G et al (2021) Human skeletal muscle mitochondrial dynamics in relation to oxidative capacity and insulin sensitivity. Diabetologia 64(2):424–436. https://doi.org/10.1007/s00125-020-05335-w
Hauffe R, Rath M, Schell M et al (2021) HSP60 reduction protects against diet-induced obesity by modulating energy metabolism in adipose tissue. Mol Metab 53:101276. https://doi.org/10.1016/j.molmet.2021.101276
Fritzen AM, Lundsgaard AM, Kiens B (2020) Tuning fatty acid oxidation in skeletal muscle with dietary fat and exercise. Nat Rev Endocrinol 16(12):683–696. https://doi.org/10.1038/s41574-020-0405-1
van Baak MA, Mariman ECM (2019) Mechanisms of weight regain after weight loss - the role of adipose tissue. Nat Rev Endocrinol 15(5):274–287. https://doi.org/10.1038/s41574-018-0148-4
Quiroga AD, Lehner R (2018) Pharmacological intervention of liver triacylglycerol lipolysis: The good, the bad and the ugly. Biochem Pharmacol 155:233–241. https://doi.org/10.1016/j.bcp.2018.07.005
Steen KA, Xu H, Bernlohr DA (2017) FABP4/aP2 regulates macrophage redox signaling and inflammasome activation via control of UCP2. Mol Cell Biol 37(2):e00282–e00216. https://doi.org/10.1128/MCB.00282-16
Gariani K, Menzies KJ, Ryu D et al (2016) Eliciting the mitochondrial unfolded protein response by nicotinamide adenine dinucleotide repletion reverses fatty liver disease in mice. Hepatology 63(4):1190–1204. https://doi.org/10.1002/hep.28245
Kumar R, Chaudhary AK, Woytash J et al (2022) A mitochondrial unfolded protein response inhibitor suppresses prostate cancer growth in mice via HSP60. J Clin Invest 132(13):e149906. https://doi.org/10.1172/JCI149906
Ansari A, Rahman MS, Saha SK, Saikot FK, Deep A, Kim KH (2017) Function of the SIRT3 mitochondrial deacetylase in cellular physiology, cancer, and neurodegenerative disease. Aging Cell 16(1):4–16. https://doi.org/10.1111/acel.12538
Ma C, Sun Y, Pi C et al (2020) Sirt3 attenuates oxidative stress damage and rescues cellular senescence in rat bone marrow mesenchymal stem cells by targeting superoxide dismutase 2. Front Cell Dev Biol 8:599376. https://doi.org/10.3389/fcell.2020.599376
Santos SS, Moreira JB, Costa M et al (2021) The mitochondrial antioxidant sirtuin3 cooperates with lipid metabolism to safeguard neurogenesis in aging and depression. Cells 11(1):90. https://doi.org/10.3390/cells11010090
Sun R, Kang X, Zhao Y et al (2020) Sirtuin 3-mediated deacetylation of acyl-CoA synthetase family member 3 by protocatechuic acid attenuates non-alcoholic fatty liver disease. Br J Pharmacol 177(18):4166–4180. https://doi.org/10.1111/bph.15159
Junli Z, Shuhan W, Yajuan Z, Xiaoling D, Jiahuan L, Keshu X (2022) The role and mechanism of CREBH regulating SIRT3 in metabolic associated fatty liver disease. Life Sci 306:120838. https://doi.org/10.1016/j.lfs.2022.120838
Zhang X, Ren X, Zhang Q et al (2016) PGC-1alpha/ERRalpha-sirt3 pathway regulates DAergic neuronal death by directly deacetylating SOD2 and ATP synthase beta. Antioxid Redox Signal 24(6):312–328. https://doi.org/10.1089/ars.2015.6403
Li Y, Wang Q, Li J, Shi B, Liu Y, Wang P (2021) SIRT3 affects mitochondrial metabolic reprogramming via the AMPK-PGC-1alpha axis in the development of benign prostatic hyperplasia. Prostate 81(15):1135–1148. https://doi.org/10.1002/pros.24208
Xin T, Lu C (2020) SirT3 activates AMPK-related mitochondrial biogenesis and ameliorates sepsis-induced myocardial injury. Aging 12(16):16224–16237. https://doi.org/10.18632/aging.103644
Kendrick AA, Choudhury M, Rahman SM et al (2011) Fatty liver is associated with reduced SIRT3 activity and mitochondrial protein hyperacetylation. Biochem J 433(3):505–514. https://doi.org/10.1042/BJ20100791
Fan F, Duan Y, Yang F et al (2020) Deletion of heat shock protein 60 in adult mouse cardiomyocytes perturbs mitochondrial protein homeostasis and causes heart failure. Cell Death Differ 27(2):587–600. https://doi.org/10.1038/s41418-019-0374-x
Acknowledgements
The authors would like to thank C.-S. Chang, C.-Y. Lin and P.-C. Tsai (Center for Mitochondrial Research and Medicine, Kaohsiung Chang Gung Memorial Hospital, Kaohsiung, Taiwan) for their invaluable technical assistance.
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Contribution statement
P-WW and S-WW developed the concept and study design. J-CW and S-WW wrote the manuscript with the support of F-SW and P-WW. J-CW and W-SL carried out the experiments. F-CS, Y-HC and Y-JS performed the data analysis. J-HC, M-HT, T-KL and C-WL contributed to the interpretation of the results. J-CW and F-SW designed the figures. M-HT, C-HS, T-HC and Y-HK provided technical support. All authors discussed the results and approved the final version. P-WW is responsible for the integrity of the work as a whole.
Funding
This work was supported by grants from Chang Gung Memorial Hospital (CMRPG8H0203 and CRRPG8K0051), the Ministry of Science and Technology, Taiwan (MOST 109-2314-B-182A-161) and the National Health Research Institutes (NHRI-EX111-11029SI).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM
(PDF 1324 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Weng, SW., Wu, JC., Shen, FC. et al. Chaperonin counteracts diet-induced non-alcoholic fatty liver disease by aiding sirtuin 3 in the control of fatty acid oxidation. Diabetologia 66, 913–930 (2023). https://doi.org/10.1007/s00125-023-05869-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-023-05869-9