Abstract
Arsenic is a major public health concern worldwide. While it is an established carcinogen and associated with a number of other adverse health outcomes, the molecular mechanisms underlying arsenic toxicity are not completely clarified. There is mounting evidence from human studies suggesting that arsenic exposure is associated with epigenetic alterations, including DNA methylation. In this review, we summarize several recent human studies that have evaluated arsenic exposure using the Illumina HumanMethylation 450K BeadChip, which interrogates more than 485,000 methylation sites across the genome. Many of these studies have observed novel regions of the genome associated with arsenic exposure. However, few studies have evaluated the biological and functional relevance of these DNA methylation changes, which are still needed. We emphasize the need for future studies to replicate the identified DNA methylation signals as well as assess whether these markers are associated with risk of arsenic-related diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
An estimated >200 million people worldwide, including ∼17 million in the USA, consume drinking water contaminated with arsenic at levels associated with a variety of adverse health effects and shortened lifespan [1, 2]. Furthermore, recent studies have suggested that food-derived inorganic arsenic exposure is also an emerging public health concern [3–7]. Epidemiologic research has established arsenic as a potent human carcinogen and has associated arsenic exposure with cancers of the skin, lung, bladder, kidney, liver, and possibly prostate [8]. Arsenic exposure has also been associated with detectably increased risk of cardiovascular disease (including coronary and ischemic heart disease, acute myocardial infarction, and hypertension) [9–11], respiratory disease [12, 13], diabetes mellitus [14–16], and impaired neurodevelopment [17]. Recent studies have observed increased infant mortality and low birth weight to be associated with prenatal arsenic exposure at moderate-to-high doses [18, 19]. With respect to its frequency, toxicity, and potential for human exposure, arsenic holds the highest ranking since 1997 on the Agency for Toxic Substances and Disease Registry (ATSDR) Substance Priority List [20].
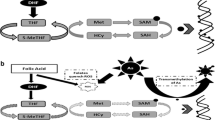
The mechanisms underlying arsenic toxicity and resulting human health effects are not completely clarified; however, epigenetic events have been hypothesized as a possible mediating pathway. There has been mounting evidence of support from animal and human studies to show that arsenic contributes to epigenetic modifications of the genome [21]. DNA methylation is an epigenetic event with a hypothesized role in gene expression, development, and disease [22]. In humans, methylation is typically of the DNA base cytosine, which is modified reversibly by adding a methyl group (–CH3) to its 5-carbon position [23]. This modification occurs on cytosines that precede a guanosine in the DNA sequence, referred to as the CpG dinucleotide. Short regions of 0.5–4 kb in length, known as CpG islands, are rich in CpG content. These islands are typically found in or near promoter regions of genes where transcription is initiated. In normal somatic cells, the vast majority of CpG dinucleotides in the genome are methylated, whereas CpG islands often remain unmethylated, allowing gene expression to occur. Whereas in disease pathways, this pattern of CpG methylation is thought to be disrupted, with increased methylation within promoter regions of genes causing abnormal gene silencing, in addition to global hypomethylation of genomic DNA, which promotes chromosomal instability, translocation, and gene disruption [24]. Unlike CpG island regions, there is greater biologic variability in methylation of CpG dinucleotides in CpG shores (within 2 kb of a CpG island), CpG shelves (2–4 kb from a CpG island), as well as isolated CpG loci in the genome [25]. DNA methylation levels are influenced by various factors including genetic, environmental, and dietary factors [26–28].
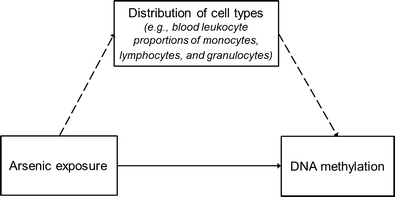
Furthermore, evidence shows cell- and tissue-specific patterns of DNA methylation, with the largest variability in DNA methylation across tissues for CpG islands located within the body of a gene as well as CpG dinucleotides located outside of CpG islands [29]. This presents an important consideration for epigenetic epidemiologic studies. First, the cellular composition of the biological specimen needs to be considered. For example, in blood-based DNA methylation studies, extracted DNA is derived from a mixture of different leukocytes (i.e., monocytes, lymphocytes, granulocytes) found in the blood. Individual cell types in blood differ not just because of specific surface markers that have been traditionally used to differentiate immune cells, but also systematically differ in DNA methylation patterns at a subset of CpG sites. Therefore, differentially methylated loci associated with an environmental factor of interest may be observed not due to a direct impact on DNA methylation but due to the effect of the environmental factor on a shift in the distribution of white blood cells, as illustrated in Fig. 1. This shift in cellular composition would therefore confound the association of interest between the environmental factor and alterations in DNA methylation. In a recently proposed novel statistical method, the DNA methylation patterns at a subset of CpG sites across the genome have been used for immunophenotyping to infer the proportions of white blood cell types in a given measured sample [30•] so that this potential source of confounding can be evaluated. Arsenic-related cell type shifts in blood have been observed in previous studies detailed below, although these cell type shifts have been estimated to account for a small proportion of the overall variability in DNA methylation. Second, the correlation of DNA methylation patterns in the measured surrogate biological tissue with the target tissue of interest needs to be considered. In human studies, access to target tissues of interest is often not feasible; therefore, studies are more commonly conducted using blood as a surrogate tissue. For the subset of CpG sites that are not correlated between leukocytes and the cells of the target tissue, differentially methylated CpG sites associated with the environmental factor in the target tissue will not be detected using DNA methylation measures from blood, missing potentially important genes and molecular pathways associated with the exposure. Conversely, differentially methylated CpG sites associated with the environmental factor in blood may be observed that are not relevant for the target tissue of interest. This is a more challenging consideration to overcome and may be informed by future publically available databases such as the Genotype-Tissue Expression Project (GTEx) as well as future studies with access to diverse tissues to enable the evaluation of tissue-specific patterns of DNA methylation.
Several human studies have examined global DNA methylation in blood in relation to arsenic exposure using surrogate markers of global DNA methylation, such as long interspersed nucleotide element-1 (LINE-1), Alu element methylation, methyl incorporation assays, or luminometric methylation assays (LUMA). The findings from these studies have been inconsistent, although they included a number of differences in exposure measures and doses across studies [31–40]. A number of studies have also evaluated arsenic in relation to candidate gene-specific DNA methylation, most frequently assessing p16 and p53 promoter methylation [36, 39, 41–45]. Studies evaluating global and candidate gene DNA methylation in relation to arsenic have been the subject of previous detailed reviews [21, 46, 47].
Epigenome-Wide DNA Methylation Studies Using Illumina 450K Platform
Relatively few epigenome-wide association studies have been conducted to investigate DNA methylation alterations in relation to arsenic exposure in humans. Published studies to date have been primarily focused on two main areas of research including the evaluation of 1) cord blood DNA methylation in relation to in utero arsenic exposure [48•, 49•, 50•, 51•] and 2) white blood cell DNA methylation in adults in relation to a variety of arsenic constructs including arsenical skin lesion status [52, 53•], urinary arsenic species [54], toenail arsenic concentration [55•], and blood arsenic and urinary total arsenic concentrations [56•]. Only one study has evaluated epigenome-wide DNA methylation patterns in tissue other than blood, which was a study of arsenic-related urothelial carcinomas [57]. Among these studies, all but three were conducted using the same epigenome-wide DNA methylation platform, the Illumina HumanMethylation 450K BeadChip (Illumina, San Diego, CA, USA), which interrogates 485,577 methylation sites per sample [48•, 49•, 50•, 51•, 53•, 55•, 56•]. Of the remaining studies, two used the Affymetrix GeneChip Human Promoter 1.0R array (Affymetrix, Santa Clara, CA, USA) [52, 54] and one used the previous generation Illumina HumanMethylation 27K BeadChip [57]. As summarized in Table 1, the studies utilizing the Illumina 450K platform are the further focus of this review.
In Utero Arsenic Exposure and Umbilical Cord Blood DNA Methylation
Four epigenome-wide association studies to date have investigated umbilical cord blood DNA methylation in relation to prenatal arsenic exposure [48•, 49•, 50•, 51•]. In a study of low-dose prenatal arsenic exposure, Koestler et al. evaluated maternal urinary arsenic concentration at 24–28 weeks gestation in relation to umbilical cord blood DNA methylation among 134 mother-child dyads from the New Hampshire Birth Cohort Study [49•]. Maternal urinary total arsenic concentration was the primary measure of in utero arsenic exposure (median (interquartile range) = 4.1 μg/L (1.8–6.6)) in the study. At a nominal significance threshold (P < 0.05), 18 % of interrogated CpG loci were associated with maternal urinary arsenic concentration, although none remained statistically significant after accounting for multiple comparisons. The authors employed a novel statistical method to infer major leukocyte components of whole blood based on a subset of DNA methylation loci [30•] and observed a significant positive association between maternal urinary total arsenic concentration and proportion of CD8+ T lymphocytes in cord blood.
A study of moderate- to high-dose prenatal arsenic exposure was conducted by Kile et al. to evaluate maternal well water arsenic concentration in relation to umbilical cord blood DNA methylation among 44 mother-child dyads from a prospective birth cohort study in Bangladesh [48•]. Maternal well water arsenic concentration at ≤16 weeks gestation was the primary measure of in utero arsenic exposure (median (range) = 12 μg/L (<1.0–581)). A positive significant association between maternal well water arsenic concentration and an intergenic DNA methylation locus on chromosome 19, cg00498691 (P = 5.89 × 10−8), was observed, after accounting for multiple comparisons. Additionally, maternal well water arsenic concentration was observed to have a significant positive association with the proportion of CD8+ T lymphocytes and a negative association with CD4+ T lymphocytes in cord blood, based on the Houseman et al. statistical method [30•].
Another study of moderate- to high-dose arsenic exposure in Bangladesh was conducted by Broberg et al. to evaluate maternal urinary total arsenic concentration in relation to umbilical cord blood DNA methylation among 127 mother-child dyads from a prenatal supplementation trial [51•]. Maternal urinary total arsenic concentrations during early (5–14 gestational weeks) and late (26–36 gestational weeks) gestation were separately evaluated as the primary measures of in utero arsenic exposure (early-gestation median (5–95 percentiles) = 66 μg/L (20–457) and late-gestation median (5–95 percentiles) = 89 μg/L (18–562)). Arsenic exposure measured in early gestation showed stronger association with DNA methylation than that in late gestation, and significant associations were only observed in boys. Three CpG sites in boys (PLIN5 cg15255455, LRRC26 cg13659051, and RPS6KA2 cg17646418) were significantly associated with early-gestation maternal urinary total arsenic concentration based on false discovery rate (FDR) <0.05. Among the top 500 nominally associated CpG loci in relation to early-gestation maternal urinary total arsenic concentration, the authors observed evidence of enrichment for hypomethylated loci among boys.
A study of moderate- to high-dose prenatal arsenic exposure was also conducted by Rojas et al. to evaluate maternal urinary total arsenic concentration in relation to umbilical cord blood DNA methylation among 38 mother-child dyads from the Biomarkers of Exposure to Arsenic (BEAR) birth cohort study in Mexico [50•]. Maternal urinary total arsenic concentration at the time of delivery was the primary measure of in utero arsenic exposure (median (range) = 32.57 μg/L (6.2–319.7)) [58]. There were 4771 differentially methylated CpG sites (34 % hypomethylated and 66 % hypermethylated) associated with maternal urinary total arsenic concentration based on FDR < 0.05. Among the arsenic-related differentially methylated CpG sites, there was evidence of enrichment for 3′UTR and gene body regions. Corresponding gene expression data were also evaluated for the 38 umbilical cord blood samples, and only weak correlations were observed for a subset of arsenic-associated CpG loci with mRNA transcript levels. The subset of CpG probes associated with gene expression changes were subsequently evaluated in relation to birth outcomes, with associations observed for gestational age, placental weight, and head circumference; the subset of genes associated with both differential DNA methylation and gene expression were also observed to be enriched for transcription factor binding sites compared to genes with altered expression but no correlation with DNA methylation.
Future Research Directions
There are notable differences across these studies making them challenging to synthesize; however, these differences also raise important issues for further exploration in future research. The studies varied with respect to the window of prenatal arsenic exposure assessment, with the study by Broberg et al. [51•] evaluating both early- and late-gestation exposures and suggesting that early prenatal exposure may have stronger effects on cord blood methylation. The studies also varied in the arsenic exposure levels of the study populations, which ranged from low- to moderate- and high-dose exposures. Additional studies will help to determine whether differentially methylated loci associated with low-dose exposure can be replicated in populations with high-dose exposure and vice versa. The study by Broberg et al. [51•] was the only analysis to present results stratified by sex, suggesting there may be important sex-specific differences in DNA methylation alterations, which should be systematically explored in future studies. Koestler et al. [49•] and Kile et al. [48•] observed an immunotoxic effect of prenatal arsenic exposure. Although the effect of arsenic on altered leukocyte cell types was estimated to explain a small percentage of the variability in methylation observed in those studies and is not believed to explain the association results observed, future studies should consider this issue as a source of potential confounding in DNA methylation analyses. Finally, a major innovation in the study by Rojas et al. [50•] was to evaluate the association of differentially methylated loci with gene expression, enrichment for transcription factor binding sites, and birth outcomes, providing additional mechanistic support through functional evidence and disease risk for the identified loci.
Arsenic and White Blood Cell DNA Methylation in Adults
Three epigenome-wide association studies have evaluated arsenic exposure in relation to white blood cell DNA methylation in adults using the Illumina 450K platform. In a prospective study of low-dose arsenic exposure, Liu et al. evaluated toenail arsenic concentration in relation to white blood cell DNA methylation 13 years later among 45 participants from the Coronary Artery Risk Development in Young Adults (CARDIA) study [55•]. Toenail arsenic concentration, a biomarker of longer-term arsenic exposure, was the main arsenic exposure measure of interest, with individuals sampled from the lowest (<0.0649 μg/g) and highest (≥0.1442 μg/g) quartiles of exposure for this study. No statistically significant associations were observed based on a strict Bonferroni P value threshold. The effect of arsenic on white blood cell proportions using the Houseman et al. statistical method [30•] was evaluated and did not reveal any significant associations with cell type proportions, although there was a modest positive association of arsenic exposure with CD8+ T lymphocyte proportions, as reported previously by others [48•, 49•].
Seow et al. conducted a prospective study among 10 incident skin lesion cases and 10 controls among adults in Bangladesh [53•]. Skin lesion cases were defined as the presence of at least one type of arsenical skin lesion. DNA methylation was measured at both baseline (2001–2003) and follow-up (2009–2011) in each study participant to identify DNA methylation changes associated with incident skin lesions based on percent methylation difference between the baseline and follow-up assessments. No differentially methylated loci were identified, after accounting for multiple comparisons, likely due to the small sample size of the study.
In the largest study to evaluate arsenic toxicity in relation to epigenome-wide DNA methylation to date, Argos et al. evaluated blood arsenic and urinary total arsenic concentrations in relation to white blood cell DNA methylation among 400 Bangladeshi adults with manifest arsenical skin lesions in cross-sectional analyses [56•]. Blood arsenic (mean = 9.3 μg/L, standard deviation (SD) = 11.3 μg/L) and urinary total arsenic (mean = 302 μg/g, SD = 364.5 μg/g) concentrations were the primary measures of arsenic exposure in the study. Statistically significant associations based on a Bonferroni significance threshold (P < 1 × 10−7) were observed for PLA2G2C cg04605617, SQSTM1 (p62) cg01225779, SLC4A4 cg06121226, and IGH cg13651690. A subset of these methylation loci were replicated in an independent study sample. Additionally, urinary total arsenic concentration was significantly inversely associated with proportions of CD4+ T lymphocytes and natural killer T cells when comparing the highest versus lowest quartiles of exposure, based on the Houseman et al. statistical method [30•]. There was evidence of enrichment for arsenic-related differentially methylated loci in ocean (isolated CpG loci in the genome) and CpG island shore regions (within 2 kb from a CpG island). The authors also evaluated the correlation of the top differentially methylated loci in relation to peripheral blood mononuclear cell gene expression and observed evidence of possible methylation-related gene regulation for a subset of these differentially methylated loci.
Future Research Directions
There are several important research considerations for future evaluations of arsenic toxicity in relation to DNA methylation alterations in adults. The study by Argos et al. [56•] was a cross-sectional analysis, whereas Liu et al. [55•] and Seow et al. [53•] took longitudinal approaches to evaluate arsenic toxicity with respect to DNA methylation alterations. Future studies should consider repeated measure analyses to evaluate the persistence of arsenic-related DNA methylation alterations. These studies all examined DNA methylation alterations in blood. The extent to which DNA methylation patterns in blood correlate with the target tissue and prove to be useful markers of disease risk or early markers of disease detection needs to be further explored [59]. Only one previous study by Yang et al. [57] evaluated epigenome-wide DNA methylation alterations in target tissue. Therefore, future studies should be designed to evaluate not only DNA methylation alterations in target organ tissues but also whether blood-based differentially methylated loci are associated with risk of arsenic-related diseases.
Research Considerations
With the emergence of epigenetic epidemiology, several research considerations are outlined for future studies evaluating arsenic epigenetics, which highlight many of the successes as well as limitations of the existing epigenome-wide DNA methylation studies evaluating arsenic toxicity.
One of the major concerns for the advancement of arsenic epigenetics and epigenetic epidemiology in general has been the interpretation of DNA methylation signals from human tissues due to heterogeneous cellular composition. This issue has been widely discussed in the literature [60]. The implications for arsenic epigenetics are such that if arsenic exposure is associated with shifts in cell types, in either blood or other tissues, then observed associations between arsenic and DNA methylation may be confounded if cellular heterogeneity is not taken into account [61]. Although arsenic-related cell type shifts in blood have been estimated to account for a small proportion of the overall variability in DNA methylation [49•], statistical methods have been proposed to account for cellular heterogeneity [30•, 62•, 63] and should be employed in future arsenic epigenetic studies to improve interpretation of study findings.
The vast majority of epigenome-wide association studies have evaluated the effect of arsenic on cord or venous blood DNA methylation. The results of blood-based studies should be interpreted cautiously since it is not established whether arsenic-related DNA methylation patterns are also present in target organ tissues. To address this, future studies should evaluate arsenic in relation to the methylome of target tissues when biological specimens are accessible. Additionally, studies examining DNA methylation alterations in white blood cells can be enhanced by evaluating differentially methylated loci in relation to arsenic-related disease outcomes in order to generate evidence on whether a differentially methylated locus identified in blood represents a biomarker of exposure or biomarker of early biological effect.
Existing epigenome-wide association studies of arsenic have been conducted in study samples with wide-ranging arsenic exposure. It is not well-established whether DNA methylation patterns observed in populations with low-dose arsenic exposure are the same as methylation patterns observed in moderate- and high-dose exposures. Additionally, future studies may shed light on the biological impacts of timing of arsenic exposure, whether differentially methylated regions vary if exposure is during in utero life, childhood, adolescence, or adulthood. Aging has an important effect on DNA methylation [64]; therefore, susceptibility of the methylome to arsenic exposure may dramatically vary by timing of arsenic exposure. Furthermore, it is not known to what extent DNA methylation alterations persist after arsenic exposure has been remediated. It will be particularly interesting for future studies to examine changes in methylation patterns among individuals with changes in arsenic exposure through prospective studies with repeated measures.
Many of the previous epigenome-wide DNA methylation studies in relation to arsenic presented findings from small study samples, which were not adequately powered to detect modest methylation effects. Studies with relatively larger sample sizes are needed to discover novel differentially methylated loci associated with arsenic as well as to replicate observed methylation signals.
Finally, epidemiologic studies are increasingly measuring multiple types of molecular or “omic” data that can be combined in order to conduct integrative molecular studies of DNA methylation and gene expression. When possible, future studies should evaluate the functional relevance of differentially methylated loci on corresponding gene expression, as was undertaken by Rojas et al. [50•] and Argos et al. [56•] to show that a subset of differentially methylated loci were associated with gene expression, which increases confidence that the identified loci might be related to disease development.
Conclusions
Epigenome-wide DNA methylation studies are a promising discovery-based approach for the identification of novel genes and pathways that may be associated with arsenic toxicity. An overview of the existing literature as well as several considerations for future studies has been presented that highlight the existing knowledge gaps in arsenic epigenetics as well as address the limitations of the existing studies. Future studies should integrate functional assessments of DNA methylation alterations as well as characterize disease risk associated with differentially methylated regions.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
IARC. http://www.inchem.org/documents/iarc/vol84/84-01-arsenic.html. 2004;84(39).
Argos M, Kalra T, Rathouz PJ, et al. Arsenic exposure from drinking water, and all-cause and chronic-disease mortalities in Bangladesh (HEALS): a prospective cohort study. Lancet. 2010;376(9737):252–8.
Gilbert-Diamond D, Cottingham KL, Gruber JF, et al. Rice consumption contributes to arsenic exposure in US women. Proc Natl Acad Sci U S A. 2011;108(51):20656–60.
Xue J, Zartarian V, Wang SW, et al. Probabilistic modeling of dietary arsenic exposure and dose and evaluation with 2003–2004 NHANES data. Environ Health Perspect. 2010;118(3):345–50.
European Food Safety Authority. EFSA Panel on Contaminants in the Food Chain (CONTAM): scientific opinion on arsenic in food. EFSA Journal. 2009;7:1351.
Potera C. U.S. rice serves up arsenic. Environ Health Perspect. 2007;115(6):A296.
Jackson BP, Taylor VF, Karagas MR, et al. Arsenic, organic foods, and brown rice syrup. Environ Health Perspect. 2012;120(5):623–6.
IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Arsenic, metals, fibres, and dusts. IARC monographs on the evaluation of carcinogenic risks to humans / World Health Organization, International Agency for Research on Cancer. 2012;100(Pt C):11–465.
Abhyankar LN, Jones MR, Guallar E, et al. Arsenic exposure and hypertension: a systematic review. Environ Health Perspect. 2012;120(4):494–500.
Navas-Acien A, Sharrett AR, Silbergeld EK, et al. Arsenic exposure and cardiovascular disease: a systematic review of the epidemiologic evidence. Am J Epidemiol. 2005;162(11):1037–49.
Moon K, Guallar E, Navas-Acien A. Arsenic exposure and cardiovascular disease: an updated systematic review. Curr Atheroscler Rep. 2012;14(6):542–55.
Mazumder DN, Steinmaus C, Bhattacharya P, et al. Bronchiectasis in persons with skin lesions resulting from arsenic in drinking water. Epidemiology. 2005;16(6):760–5.
Parvez F, Chen Y, Yunus M, et al. Arsenic exposure and impaired lung function. Findings from a large population-based prospective cohort study. Am J Respir Crit Care Med. 2013;188(7):813–9.
Maull EA, Ahsan H, Edwards J, et al. Evaluation of the association between arsenic and diabetes: a national toxicology program workshop review. Environ Health Perspect. 2012. doi:10.1289/ehp.1104579.
Thayer KA, Heindel JJ, Bucher JR, et al. Role of environmental chemicals in diabetes and obesity: a National Toxicology Program workshop review. Environ Health Perspect. 2012;120(6):779–89.
Navas-Acien A, Silbergeld EK, Streeter RA, et al. Arsenic exposure and type 2 diabetes: a systematic review of the experimental and epidemiological evidence. Environ Health Perspect. 2006;114(5):641–8.
Rodriguez-Barranco M, Lacasana M, Aguilar-Garduno C, et al. Association of arsenic, cadmium and manganese exposure with neurodevelopment and behavioural disorders in children: a systematic review and meta-analysis. Sci Total Environ. 2013;454–455:562–77.
Rahman A, Vahter M, Ekstrom EC, et al. Association of arsenic exposure during pregnancy with fetal loss and infant death: a cohort study in Bangladesh. Am J Epidemiol. 2007;165(12):1389–96.
Huyck KL, Kile ML, Mahiuddin G, et al. Maternal arsenic exposure associated with low birth weight in Bangladesh. J Occup Environ Med. 2007;49(10):1097–104.
Agency for Toxic Substances and Disease Registry. The ATSDR 2011 substance priority list 2011 [cited 2013 May 31]. Available from: http://www.atsdr.cdc.gov/SPL/.
Bailey KA, Fry RC. Arsenic-associated changes to the epigenome: what are the functional consequences? Curr Environ Health Rep. 2014;1:22–34.
El-Osta A, Wolffe AP. DNA methylation and histone deacetylation in the control of gene expression: basic biochemistry to human development and disease. Gene Expr. 2000;9(1–2):63–75.
Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med. 2003;349(21):2042–54.
Esteller M. Cancer epigenomics: DNA methylomes and histone-modification maps. Nat Rev. 2007;8(4):286–98.
Ziller MJ, Gu H, Muller F, et al. Charting a dynamic DNA methylation landscape of the human genome. Nature. 2013;500(7463):477–81.
McKay JA, Groom A, Potter C, et al. Genetic and non-genetic influences during pregnancy on infant global and site specific DNA methylation: role for folate gene variants and vitamin B(12). PLoS ONE. 2012;7(3):e33290.
Siedlinski M, Klanderman B, Sandhaus RA, et al. Association of cigarette smoking and CRP levels with DNA methylation in alpha-1 antitrypsin deficiency. Epigenetics. 2012;7(7).
Philibert RA, Plume JM, Gibbons FX, et al. The impact of recent alcohol use on genome wide DNA methylation signatures. Front Genet. 2012;3:54.
Lokk K, Modhukur V, Rajashekar B, et al. DNA methylome profiling of human tissues identifies global and tissue-specific methylation patterns. Genome Biol. 2014;15(4):r54.
Houseman EA, Accomando WP, Koestler DC, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics. 2012;13:86. This paper presents a novel statistical method for inferring leukocytes proportions based on a set of informative DNA methylation markers.
Pilsner JR, Liu X, Ahsan H, et al. Genomic methylation of peripheral blood leukocyte DNA: influences of arsenic and folate in Bangladeshi adults. Am J Clin Nutr. 2007;86(4):1179–86.
Pilsner JR, Liu X, Ahsan H, et al. Folate deficiency, hyperhomocysteinemia, low urinary creatinine, and hypomethylation of leukocyte DNA are risk factors for arsenic-induced skin lesions. Environ Health Perspect. 2009;117(2):254–60.
Majumdar S, Chanda S, Ganguli B, et al. Arsenic exposure induces genomic hypermethylation. Environ Toxicol. 2010;25(3):315–8.
Wilhelm CS, Kelsey KT, Butler R, et al. Implications of LINE1 methylation for bladder cancer risk in women. Clin Cancer Res. 2010;16(5):1682–9.
Lambrou A, Baccarelli A, Wright RO, et al. Arsenic exposure and DNA methylation among elderly men. Epidemiology. 2012;23(5):668–76.
Hossain MB, Vahter M, Concha G, et al. Environmental arsenic exposure and DNA methylation of the tumor suppressor gene p16 and the DNA repair gene MLH1: effect of arsenic metabolism and genotype. Metallomics. 2012;4(11):1167–75.
Tajuddin SM, Amaral AF, Fernandez AF, et al. Genetic and non-genetic predictors of LINE-1 methylation in leukocyte DNA. Environ Health Perspect. 2013. doi:10.1289/ehp.1206068.
Kile ML, Baccarelli A, Hoffman E, et al. Prenatal arsenic exposure and DNA methylation in maternal and umbilical cord blood leukocytes. Environ Health Perspect. 2012;120(7):1061–6.
Intarasunanont P, Navasumrit P, Waraprasit S, et al. Effects of arsenic exposure on DNA methylation in cord blood samples from newborn babies and in a human lymphoblast cell line. Environ Heal. 2012;11:31.
Pilsner JR, Hall MN, Liu X, et al. Influence of prenatal arsenic exposure and newborn sex on global methylation of cord blood DNA. PLoS ONE. 2012;7(5):e37147.
Chen WT, Hung WC, Kang WY, et al. Urothelial carcinomas arising in arsenic-contaminated areas are associated with hypermethylation of the gene promoter of the death-associated protein kinase. Histopathology. 2007;51(6):785–92.
Zhang AH, Bin HH, Pan XL, et al. Analysis of p16 gene mutation, deletion and methylation in patients with arseniasis produced by indoor unventilated-stove coal usage in Guizhou, China. J Toxicol Environ Health. 2007;70(11):970–5.
Marsit CJ, Karagas MR, Danaee H, et al. Carcinogen exposure and gene promoter hypermethylation in bladder cancer. Carcinogenesis. 2006;27(1):112–6.
Chanda S, Dasgupta UB, Guhamazumder D, et al. DNA hypermethylation of promoter of gene p53 and p16 in arsenic-exposed people with and without malignancy. Toxicol Sci. 2006;89(2):431–7.
Engstrom KS, Hossain MB, Lauss M, et al. Efficient arsenic metabolism–the AS3MT haplotype is associated with DNA methylation and expression of multiple genes around AS3MT. PLoS ONE. 2013;8(1):e53732.
Ren X, McHale CM, Skibola CF, et al. An emerging role for epigenetic dysregulation in arsenic toxicity and carcinogenesis. Environ Health Perspect. 2011;119(1):11–9.
Reichard JF, Puga A. Effects of arsenic exposure on DNA methylation and epigenetic gene regulation. Epigenomics. 2010;2(1):87–104.
Kile ML, Houseman EA, Baccarelli A, et al. Effect of prenatal arsenic exposure on DNA methylation and leukocyte subpopulations in cord blood. Epigenetics. 2014;9(5). This study observed an association between prenatal arsenic exposure and cord blood DNA methylation, adjusting analyses for cell heterogeneity
Koestler DC, Avissar-Whiting M, Houseman EA, et al. Differential DNA methylation in umbilical cord blood of infants exposed to low levels of arsenic in utero. Environ Health Perspect. 2013;121(8):971–7. This study evaluated prenatal arsenic exposure on methylomic changes and is particularly informative for the analyses evaluating the genomic location as well as effect of arsenic on cell heterogeneity.
Rojas D, Rager JE, Smeester L, et al. Prenatal arsenic exposure and the epigenome: identifying sites of 5-methyl cytosine alterations that predict functional changes in gene expression in newborn cord blood and subsequent birth outcomes. Toxicol Sci. 2014. doi:10.1093/toxsci/kfu210. This study observed associations between prenatal arsenic exposure and cord blood DNA methylation, as well as evaluated functional evidence using gene expression data, and validation through disease outcomes.
Broberg K, Ahmed S, Engstrom K, et al. Arsenic exposure in early pregnancy alters genome-wide DNA methylation in cord blood, particularly in boys. J Dev Orig Health Dis. 2014;5(4):288–98. This study observed associations between prenatal arsenic exposure and cord blood DNA methylation among boys, showing sex-specific effects of arsenic on the methylome.
Smeester L, Rager JE, Bailey KA, et al. Epigenetic changes in individuals with arsenicosis. Chem Res Toxicol. 2011;24(2):165–7.
Seow WJ, Kile ML, Baccarelli AA, et al. Epigenome-wide DNA methylation changes with development of arsenic-induced skin lesions in Bangladesh: a case–control follow-up study. Environ Mol Mutagen. 2014;55(6):449–56. Although no significant associations were observed, this study employed a repeated assessment of DNA methylation to evaluate changes in methylation in relation to skin lesion incidence.
Bailey KA, Wu MC, Ward WO, et al. Arsenic and the epigenome: interindividual differences in arsenic metabolism related to distinct patterns of DNA methylation. J Biochem Mol Toxicol. 2013;27(2):106–15.
Liu X, Zheng Y, Zhang W, et al. Blood methylomics in response to arsenic exposure in a low-exposed US population. J Expo Sci Environ Epidemiol. 2014;24(2):145–9. Although no significant associations were observed, this study conducted a prospective analysis of arsenic exposure on subsequent DNA methylation alterations.
Argos M, Chen L, Jasmine F, et al. Gene-specific differential DNA methylation and chronic arsenic exposure in an epigenome-wide association study of adults in Bangladesh. Environ Health Perspect. 2015;123(1):64–71. This study observed associations between blood and urinary arsenic exposure and white blood cell DNA methylation, evaluated functional evidence using gene expression data, and replicated a subset of study findings in an independent cohort.
Yang TY, Hsu LI, Chiu AW, et al. Comparison of genome-wide DNA methylation in urothelial carcinomas of patients with and without arsenic exposure. Environ Res. 2014;128:57–63.
Rager JE, Bailey KA, Smeester L, et al. Prenatal arsenic exposure and the epigenome: altered microRNAs associated with innate and adaptive immune signaling in newborn cord blood. Environ Mol Mutagen. 2014;55(3):196–208.
Talens RP, Boomsma DI, Tobi EW, et al. Variation, patterns, and temporal stability of DNA methylation: considerations for epigenetic epidemiology. FASEB J. 2010;24(9):3135–44.
Liang L, Cookson WO. Grasping nettles: cellular heterogeneity and other confounders in epigenome-wide association studies. Hum Mol Genet. 2014;23(R1):R83–8.
Reinius LE, Acevedo N, Joerink M, et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS ONE. 2012;7(7):e41361.
Houseman EA, Molitor J, Marsit CJ. Reference-free cell mixture adjustments in analysis of DNA methylation data. Bioinformatics. 2014;30(10):1431–9. This paper presents a novel statistical method for inferring cell type proportions for tissues (e.g., placenta, saliva, adipose or tumor tissue) where the relevant underlying cell types may not be known.
Winham SJ, Armasu SM, Cicek MS, et al. Genome-wide investigation of regional blood-based DNA methylation adjusted for complete blood counts implicates BNC2 in ovarian cancer. Genet Epidemiol. 2014;38(5):457–66.
Bjornsson HT, Sigurdsson MI, Fallin MD, et al. Intra-individual change over time in DNA methylation with familial clustering. JAMA. 2008;299(24):2877–83.
Acknowledgments
Maria Argos reports grants from NIH.
Compliance with Ethics Guidelines
ᅟ
Conflict of Interest
Maria Argos declares that she has no conflict of interest.
Human and Animal Rights and Informed Consent
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection on Environmental Epigenetics
Rights and permissions
About this article
Cite this article
Argos, M. Arsenic Exposure and Epigenetic Alterations: Recent Findings Based on the Illumina 450K DNA Methylation Array. Curr Envir Health Rpt 2, 137–144 (2015). https://doi.org/10.1007/s40572-015-0052-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40572-015-0052-1